GTP vs GDP: Structural Secrets of Microtubule Dynamics and Their Impact on Drug Discovery
This article provides a comprehensive comparison of GTP- and GDP-bound microtubule structures, targeting researchers and drug development professionals.
GTP vs GDP: Structural Secrets of Microtubule Dynamics and Their Impact on Drug Discovery
Abstract
This article provides a comprehensive comparison of GTP- and GDP-bound microtubule structures, targeting researchers and drug development professionals. It explores the foundational biochemistry of tubulin's GTPase activity and its role in dynamic instability. The piece details advanced methodological approaches for structural elucidation, addresses common experimental challenges in studying these transient states, and validates key structural differences through comparative analysis. The synthesis offers crucial insights for targeting microtubules in cancer therapy and neurodegenerative diseases.
The GTPase Engine: Understanding the Biochemical Basis of Microtubule Dynamics
The structural integrity and dynamic behavior of microtubules are fundamentally governed by the properties of their α/β-tubulin heterodimeric building blocks. A critical, yet often underemphasized, site of control is the nucleotide pocket on β-tubulin, termed the Exchangeable or E-site. This guide provides a comparative analysis of microtubule performance based on the nucleotide state (GTP vs. GDP) in this pocket, a central thesis in understanding microtubule dynamics and stability.
Comparison Guide: GTP- vs. GDP-β-Tubulin in Microtubule Assembly & Stability
The nucleotide bound at the E-site of β-tubulin dictates the conformation of the dimer and its interactions within the polymer. The following table summarizes key performance metrics derived from in vitro reconstitution experiments.
Table 1: Comparative Performance of GTP- vs. GDP-Microtubules
| Performance Metric | GTP-Microtubule (GTP-Cap State) | GDP-Microtubule (Lattice Core) | Experimental Support & Notes |
|---|---|---|---|
| Polymerization Rate | High (Fast elongation) | Not Applicable (Stable lattice) | Measured by turbidity (A350) or TIRF microscopy. GTP-state promotes favorable lateral contacts. |
| Critical Concentration (Cc) | Low (~2-4 µM for pure tubulin) | Very High (>20 µM) | Cc is the tubulin concentration at which assembly begins. GTP-form is polymerization-competent. |
| Microtubule Stability | Dynamic (Prone to depolymerization if GTP hydrolyzes) | Low (GDP-lattice is intrinsically curved and unstable) | Basis for "dynamic instability." GDP-tubulin favors a curved conformation incompatible with the straight polymer. |
| Lateral Interaction Strength | Strong | Weak | Cryo-EM shows tighter interfacial bonds in GTP-like structures. GDP-state weakens dimer-dimer contacts. |
| Protofilament Curvature | Straight (within polymer) | Curved (~12° angle from longitudinal axis) | Visualized by cryo-EM of depolymerizing ends or GDP-tubulin rings. |
| Drug Susceptibility (e.g., Taxol) | Binds, stabilizes | Binding enhances, but lattice is inherently less stable | Taxol primarily binds and stabilizes the GDP-lattice, suppressing catastrophe. |
Experimental Protocols for Key Comparisons
1. Measuring Polymerization Kinetics & Critical Concentration
- Protocol: Purified tubulin is clarified by centrifugation at 4°C to remove aggregates. Assembly is initiated by rapidly shifting to 37°C in BRB80 buffer (80 mM PIPES, 1 mM MgCl₂, 1 mM EGTA, pH 6.9) with 1 mM GTP. Polymerization is monitored by turbidity at 350 nm in a spectrophotometer or via TIRF microscopy using rhodamine-labeled tubulin. To determine the Critical Concentration (Cc), the final plateau absorbance (or polymer mass) from multiple reactions at different tubulin concentrations is plotted against the total tubulin concentration. The x-intercept is the Cc.
- Data Interpretation: A low Cc indicates a highly assembly-competent system (characteristic of GTP-state). The lag phase, growth rate, and plateau provide data on nucleation and elongation.
2. Visualizing Dynamic Instability & GTP-Cap Behavior
- Protocol: Microtubules are polymerized from stabilized seeds (e.g., GMPCPP seeds) in a flow chamber. Elongation occurs in the presence of a low concentration of labeled tubulin (e.g., 10-15 µM total tubulin, <5% labeled) and an oxygen-scavenging system for TIRF microscopy. Growth and shrinkage events are tracked and quantified over time.
- Data Interpretation: Microtubules with a protective GTP-cap exhibit prolonged growth. Stochastic loss of the cap (through hydrolysis) leads to a "catastrophe" and rapid depolymerization of the GDP-core, followed by "rescue" and regrowth.
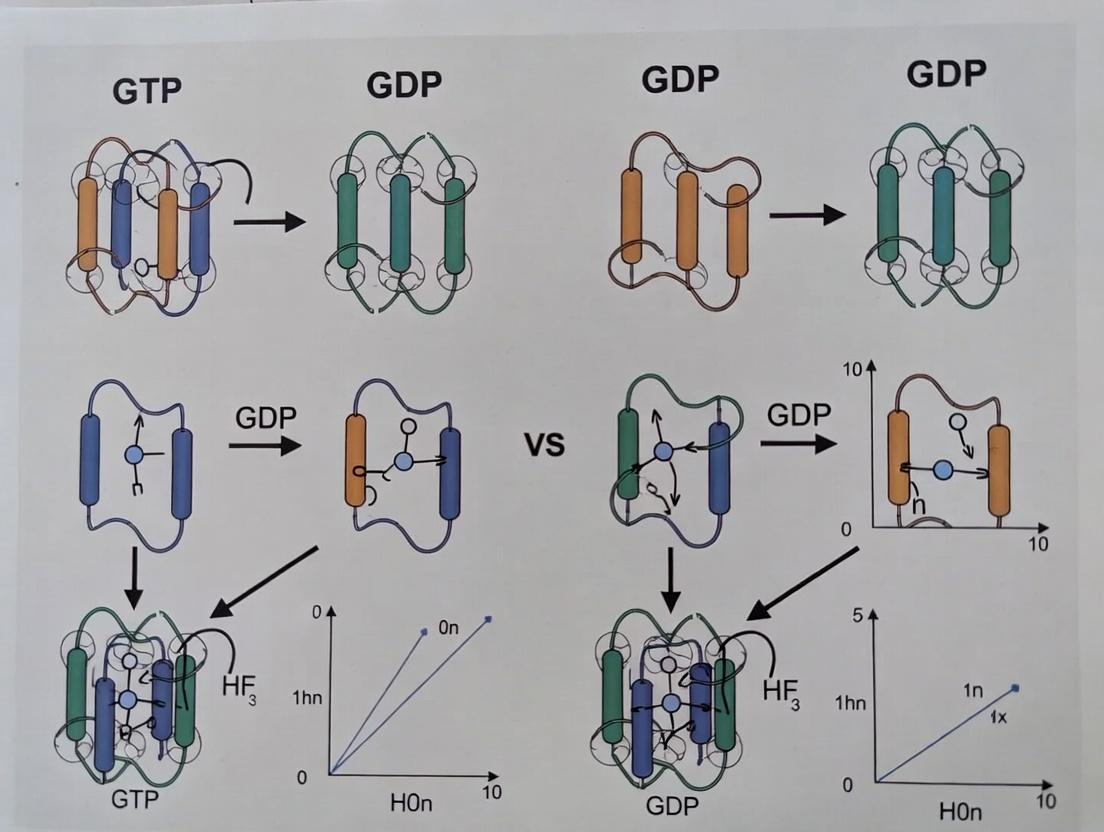
Visualization of Tubulin States & Microtubule Dynamics
Diagram Title: GTP-Cap Model of Microtubule Dynamic Instability
The Scientist's Toolkit: Key Research Reagents & Materials
Table 2: Essential Reagents for Tubulin Nucleotide-State Research
| Reagent/Material | Function & Role in Comparison |
|---|---|
| Purified Tubulin (>99%) | The fundamental substrate. Must be nucleotide-free or precisely loaded for clean experiments. |
| GTP (Guanosine-5'-triphosphate) | The natural exchangeable nucleotide. Required for polymerization-competent dimers. |
| GDP (Guanosine-5'-diphosphate) | The hydrolysis product. Used to pre-form GDP-dimers to study polymerization incompetence. |
| GMPCPP (Guanylyl-(α,β)-methylene-diphosphonate) | A non-hydrolyzable GTP analog. Creates permanently stable "GTP-like" microtubules for structural studies. |
| Taxol/Paclitaxel | A small molecule that binds and stabilizes the GDP-lattice, suppressing dynamic instability. |
| BRB80 Buffer | Standard physiological buffer for microtubule experiments, maintains pH and cation conditions. |
| TIRF Microscope | Enables real-time, single-microtubule visualization of growth and shrinkage dynamics. |
| Cryo-Electron Microscopy (Cryo-EM) | Provides high-resolution 3D structures of tubulin dimers and microtubules in different nucleotide states. |
Within the broader research thesis comparing GTP- versus GDP-microtubule structures, the GTP Cap Hypothesis remains the central model explaining microtubule dynamic instability. This phenomenon, characterized by stochastic growth and rapid shortening, is governed by the nucleotide state of tubulin subunits at the microtubule end. This guide compares the core tenets and supporting experimental evidence for the GTP Cap Hypothesis against alternative theoretical models.
Model Comparison Guide
Table 1: Comparison of Models Explaining Microtubule Dynamic Instability
| Model / Hypothesis | Core Premise | Key Predictive Differences | Experimental Support Status |
|---|---|---|---|
| GTP Cap Hypothesis | A stabilizing "cap" of GTP-tubulin at the microtubule end prevents catastrophic depolymerization; hydrolysis to GDP-tubulin in the lattice creates strain, leading to rapid shrinkage if the cap is lost. | Catastrophe frequency depends on cap size/stability. Rescue requires re-forming a GTP cap. | Strong; supported by direct and indirect evidence from kinetics, mutant studies, and analogs. |
| Conformational Switch Model | The nucleotide state (GTP vs. GDP) induces a conformational change in tubulin, altering its lateral bonding strength within the lattice. | Dynamics are driven by lattice strain and bond geometry, not solely by a protective cap. | Complementary; seen as a mechanistic detail of the cap model. |
| Stochastic Model | Dynamic instability can be explained by random tubulin addition/loss without requiring a structural cap, based on kinetics of a two-state system. | Predicts relationships between growth rate, catastrophe frequency, and dilution experiments. | Partially supported; but does not fully explain all kinetic data without incorporating a cap concept. |
| Lattice Strain Model | Focuses on the mechanical strain stored in the GDP-lattice as the primary driver of catastrophe, with the cap merely delaying its release. | Emphasizes the role of lattice geometry and the "power stroke" during shrinkage. | Integrated; now considered part of the modern synthesis of the cap hypothesis. |
Supporting Experimental Data & Protocols
Key Experiment 1: Kinetic Analysis of Microtubule Growth with Non-Hydrolyzable GTP Analogs
Protocol:
- Purify tubulin free of endogenous nucleotides via ion-exchange chromatography.
- Prepare elongation buffer containing a non-hydrolyzable GTP analog (e.g., GMPCPP or GTPγS).
- Use differential interference contrast (DIC) or TIRF microscopy to visualize microtubules nucleated from stabilized seeds.
- Initiate growth by introducing analog-bound tubulin to the chamber.
- Measure elongation rates and observe filament stability over time.
Table 2: Microtubule Growth Parameters with Different Nucleotides
| Nucleotide Condition | Average Elongation Rate (µm/min) | Catastrophe Frequency (events/min) | Average Shrinkage Rate (µm/min) | Observed Outcome |
|---|---|---|---|---|
| GMPCPP (Analog) | 5.2 ± 1.1 | ~0 | N/A | Stable, non-dynamic polymers. No catastrophes observed. |
| GTP (Standard) | 12.8 ± 2.3 | 0.05 ± 0.02 | 24.5 ± 3.8 | Normal dynamic instability. |
| GDP (Control) | N/A | Immediate | 28.1 ± 4.2 | No growth; pure depolymerization. |
Interpretation: The non-hydrolyzable analog forms a permanent "GTP cap," resulting in perfectly stable microtubules, directly supporting the hypothesis that GTP hydrolysis is necessary for catastrophe.
Key Experiment 2: Cap Size Estimation via Dilution-Induced Catastrophe
Protocol:
- Grow microtubules from centrosomal seeds or GMPCPP seeds in a flow chamber.
- Monitor individual microtubules during steady-state growth in a known tubulin concentration (e.g., 12 µM).
- Rapidly flush the chamber with tubulin-free buffer (dilution experiment).
- Record the time lag between dilution and the onset of catastrophe for a population of microtubules.
- Calculate the cap size based on the growth rate and the mean time to catastrophe post-dilution.
Data: The time lag implies a protective cap of approximately 100-200 tubulin dimers at the growing end under these conditions.
Visualization of Core Concepts
Diagram Title: GTP Cap Maintenance vs. Loss Leading to Catastrophe
Diagram Title: Key Experimental Workflow for Testing the GTP Cap
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for GTP Cap & Dynamic Instability Research
| Reagent / Material | Function & Relevance to Hypothesis |
|---|---|
| Purified Tubulin (>99% pure) | Essential substrate. Must be free of contaminating nucleotides for clean kinetic studies and analog incorporation. |
| Non-Hydrolyzable GTP Analogs (GMPCPP, GTPγS) | Form permanent GTP caps, proving cap function. Used to create stable microtubule seeds for assays. |
| Fluorescently-Labeled Tubulin (e.g., HiLyte 488, TAMRA) | Enables real-time visualization of microtubule dynamics via TIRF microscopy. |
| Taxol (Paclitaxel) & GTPase-Deficient Tubulin Mutants | Positive controls that stabilize microtubules, mimicking a permanent cap. |
| Nocodazole / Colchicine | Negative controls that promote depolymerization, used in dilution/chase experiments. |
| TIRF Microscopy System | High-resolution, single-filament imaging required to measure growth/shrinkage rates and catastrophe events. |
| Enzymatic Assay Kits (e.g., Malachite Green Phosphate) | Quantify GTP hydrolysis rates of free tubulin vs. polymerized microtubules. |
This guide provides a comparative analysis of the functional "products" in microtubule dynamics: GTP-bound and GDP-bound tubulin dimers. Framed within broader research on microtubule structural plasticity, we evaluate their performance based on stability, lattice geometry, and interaction interfaces.
Conformational and Functional Comparison
The following table summarizes key quantitative differences between the two states, derived from structural and biophysical studies.
Table 1: Comparative Properties of GTP- vs. GDP-Tubulin Dimers
| Property | GTP-Tubulin (Straight) | GDP-Tubulin (Kinked) | Experimental Method (Typical) |
|---|---|---|---|
| Intra-dimer Curvature | ~0° (Straight) | ~12° - 22° (Kinked) | High-Resolution Cryo-EM |
| Inter-dimer Longitudinal Bond Strength | Strong | Weakened (~1000-fold reduction) | Kinetic Measurements, Optical Traps |
| Microtubule Lattice Incorporation | Preferred (Stabilizing) | Disfavored (Destabilizing) | In Vitro Polymerization Assays |
| Lateral Contact Interface | Compact, Complementary | Disrupted, Weakened | X-ray Crystallography, Molecular Dynamics |
| Susceptibility to Depolymerization | Low (Protected 'GTP-cap') | High (Core of shrinking MT) | Turbidity Assays, TIRF Microscopy |
Detailed Experimental Protocols
1. Cryo-EM for Determining Tubulin Conformation
- Objective: Solve high-resolution structures of tubulin in different nucleotide states within microtubules or as isolated dimers.
- Protocol: Tubulin is polymerized in the presence of non-hydrolyzable GTP analogs (e.g., GMPCPP) to stabilize GTP-like states, or with GDP for the GDP-state. Samples are vitrified on EM grids. Data is collected on a cryo-electron microscope, followed by single-particle analysis or helical reconstruction to generate 3D density maps. Dimer curvature and lateral contact interfaces are measured from the refined atomic models.
2. Microtubule Dynamic Assay (TIRF Microscopy)
- Objective: Visualize and quantify the differential incorporation and stability of GTP- vs. GDP-tubulin.
- Protocol: Biotinylated GMPCPP (GTP-like) microtubule seeds are immobilized on a passivated glass slide in a flow chamber. Unlabeled tubulin at a critical concentration is flowed in with a small fraction of fluorescently labeled tubulin (Hilyte 488 or similar) and an oxygen scavenging system for imaging. Growth and shrinkage events at both ends are tracked over time using Total Internal Reflection Fluorescence (TIRF) microscopy to determine catastrophe and rescue frequencies, directly reporting on the stability conferred by the GTP-cap versus the GDP-core.
3. Kinetic Analysis of Tubulin Dimer Affinity
- Objective: Measure the strength of longitudinal interactions between dimers in different nucleotide states.
- Protocol: Using surface plasmon resonance (SPR) or isothermal titration calorimetry (ITC), one tubulin dimer (the ligand) is immobilized or titrated into a solution containing another (the analyte). The association (
k_on) and dissociation (k_off) rate constants, as well as the equilibrium dissociation constant (K_D), are measured for combinations involving GMPCPP (GTP-like) or GDP-bound tubulin, quantifying the nucleotide-dependent bond strength.
Visualization of Concepts
Title: GTP Hydrolysis Drives Conformational and Stability Switch
Title: Cryo-EM Workflow for Tubulin Structure
The Scientist's Toolkit: Key Research Reagents
Table 2: Essential Reagents for GTP/GDP-Tubulin Studies
| Reagent | Function & Rationale |
|---|---|
| Non-hydrolyzable GTP Analogs (GMPCPP, GTPγS) | Mimics the GTP-bound state indefinitely, allowing study of stable, straight microtubules and GTP-like tubulin conformation without hydrolysis. |
| Taxol/Paclitaxel | Binds and stabilizes the microtubule lattice, often used to study GDP-tubulin in a polymerized context by suppressing depolymerization. |
| Biotinylated Tubulin & NeutrAvidin | Enables surface immobilization of microtubule seeds for TIRF microscopy-based dynamic assays. |
| Oxygen Scavenging System (e.g., PCA/PCD) | Reduces photobleaching and radical damage during prolonged fluorescence microscopy (e.g., TIRF), allowing longer observation times. |
| Cysteine-reactive Fluorescent Dyes (e.g., Cy3, Alexa Fluor 488-maleimide) | Site-specific labeling of engineered tubulin cysteine residues for tracking dimer incorporation in kinetic or imaging experiments. |
| Tubulin Purification Kits (from bovine/porcine brain or recombinant) | Provides high-purity, functional tubulin, the essential substrate for all in vitro biochemical and structural studies. |
This guide compares the performance of key methodologies and tools used to investigate the GTP to GDP conversion trigger in microtubules, a critical process for understanding dynamic instability and a central focus of GTP vs GDP microtubule structure comparison research.
Comparison of Key Experimental Approaches
Table 1: Comparison of Hydrolysis Rate Measurement Techniques
| Method | Principle | Temporal Resolution | Spatial Information | Key Advantage | Key Limitation | Typical k_hyd (s⁻¹) Measurement Range* |
|---|---|---|---|---|---|---|
| Cap Expansion Assay | Measures growth of stable GDP-microtubule seed after GTP-tubulin addition. | Seconds to minutes. | Low (bulk). | Simple, measures functional outcome in ensemble. | Indirect, assumes hydrolysis is rate-limiting. | 0.05 - 0.5 |
| FRET-Based Probes | Uses labeled tubulin with donor/acceptor to report conformational change post-hydrolysis. | Millisecond to second. | Low to medium (can be single MT). | Direct, reports on chemical step or structural change. | Probe labeling may alter kinetics. | 0.1 - 10 |
| Cryo-EM & Time-Resolved Analysis | Traps intermediates at defined time points for high-resolution structure determination. | Milliseconds (with rapid mixing/freezing). | Atomic (single MT). | Direct structural mechanism; "sees" the trigger. | Technically challenging; not real-time in solution. | N/A (Structural) |
| MD Simulations | Computationally models atomic interactions and energy landscapes over time. | Femtoseconds to microseconds. | Atomic. | Proposes testable atomic-level mechanisms. | Computationally limited; force field dependent. | N/A (Theoretical) |
| Mutant Tubulin Analysis | Measures kinetics of hydrolysis-deficient (e.g., Q61H β-tubulin) or altering mutants. | Seconds to minutes. | Low (bulk). | Identifies critical residues for the trigger. | Mutations may cause pleiotropic effects. | Varies by mutant |
*Reported hydrolysis rates for unperturbed microtubules vary, with typical values near ~0.5 s⁻¹ at the plus end.
Detailed Experimental Protocols
Protocol 1: Cap Expansion Assay for Ensemble Hydrolysis Rate Estimation
Objective: Infer the GTP hydrolysis rate constant from the kinetics of GDP-microtubule seed elongation.
- Seed Preparation: Stabilize GMPCPP (non-hydrolyzable analog) microtubule seeds onto a coverslip in a flow chamber.
- Initial Growth: Flush in a solution of purified tubulin (10-15 µM) in BRB80 buffer (80 mM PIPES, 1 mM MgCl₂, 1 mM EGTA, pH 6.8) with 1 mM GTP. Incubate 5-10 min to form a GTP-cap.
- Trigger Dilution: Rapidly dilute the tubulin concentration 10-50x with pre-warmed BRB80 + GTP buffer. This halts new tubulin addition, isolating cap hydrolysis and depolymerization.
- Imaging & Analysis: Acquire time-lapse images via TIRF microscopy. Measure the length of the stable GDP-microtubule (the seed + the post-dilution grown segment) over time. The initial slope of growth post-dilution, before catastrophe, approximates the hydrolysis rate.
Protocol 2: FRET-Based Kinetics Measurement with 2'(3')-O-(N-Methylanthraniloyl) (Mant)-GTP
Objective: Directly monitor the chemical step of GTP hydrolysis on microtubules in real-time.
- Probe Preparation: Pre-load purified tubulin (40 µM) with mant-GTP (100 µM) in polymerization buffer on ice for 30 min.
- Baseline Acquisition: In a fluorometer cuvette, place mant-GTP-loaded tubulin in polymerization buffer at 10°C (to inhibit polymerization). Excite at 355 nm, record emission at 440 nm (mant signal).
- Reaction Initiation: Rapidly shift temperature to 37°C to initiate microtubule polymerization and hydrolysis.
- Data Acquisition: Record the fluorescence decrease over time (seconds to minutes) as mant-GTP converts to mant-GDP. Fit the decay curve to a single exponential to obtain the observed rate constant.
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for GTP Hydrolysis Studies
| Item | Function & Rationale |
|---|---|
| Non-hydrolyzable GTP Analogs (GMPCPP, GMPPNP) | Form stable, non-dynamic microtubules; serve as structural and experimental controls to isolate hydrolysis effects. |
| Fluorescent GTP Analogs (Mant-GTP, BODIPY-GTP) | Enable real-time spectroscopic (FRET, direct fluorescence) monitoring of nucleotide state change in solution or on microtubules. |
| Hydrolysis-Deficient Mutant Tubulins (e.g., β-tubulin Q61H) | Allow dissection of the role of specific residues in the hydrolysis trigger and separate hydrolysis from polymerization effects. |
| Caged GTP | Enables precise, UV light-triggered initiation of polymerization and hydrolysis for ultra-fast kinetic studies (millisecond resolution). |
| Stabilizing Agents (Taxol, Zampanolide) | Lock microtubules in a specific state (largely GDP-like) to study hydrolysis intermediates or separate dynamics from stability. |
| Kinesin Motors (e.g., Kif5b) | Used as structural probes, as their binding is sensitive to microtubule nucleotide state, reporting on GTP cap size and hydrolysis timing. |
Visualization of Key Concepts
Title: GTP Hydrolysis Pathway in Microtubule Dynamic Instability
Title: Integrated Workflow for Studying the Hydrolysis Trigger
Within the broader thesis on GTP vs. GDP microtubule structure comparison, a critical distinction lies in the location and function of the guanine nucleotide within the tubulin heterodimer. This guide provides an objective comparison of the roles of GTP bound at the exchangeable site (E-site) and the non-exchangeable site (N-site) in microtubule dynamics and stability, supported by current experimental data.
Structural and Functional Comparison
The Two GTP-Binding Sites:
- E-site (Exchangeable): Located on β-tubulin. This site can exchange GTP for GDP and is hydrolyzed to GDP during or shortly after polymerization. The nucleotide state at this site controls the stability of the microtubule lattice.
- N-site (Non-exchangeable): Located on α-tubulin. This site is permanently bound by GTP (or a non-hydrolyzable analog) and is never hydrolyzed under physiological conditions. It plays a structural role in dimer formation and longitudinal contacts.
The table below summarizes the core characteristics and roles of each site.
Table 1: Core Characteristics of E-site and N-site GTP
| Feature | E-site (β-tubulin) | N-site (α-tubulin) |
|---|---|---|
| Nucleotide Exchange | Exchangeable (GTP ⇄ GDP) | Non-exchangeable (permanently GTP-bound) |
| Hydrolysis | Hydrolyzed to GDP post-incorporation | Not hydrolyzed |
| Primary Role | Provides energy for dynamic instability; creates a "GTP-cap" for stability | Structural; essential for heterodimer formation and longitudinal interface integrity |
| Consequence of State Change | GDP in lattice leads to lattice strain and catastrophic depolymerization | Loss of GTP binding disrupts dimerization, preventing polymerization |
| Drug Targeting | Major site for anti-mitotic agents (e.g., Taxol stabilizes GDP-lattice) | Not a direct drug target due to buried, non-exchangeable nature |
Quantitative Data on Functional Impact
Experimental data from kinetic studies, cryo-EM reconstructions, and lattice stability assays highlight the distinct contributions of each site.
Table 2: Experimental Data on Polymerization and Stability Parameters
| Parameter & Measurement Method | E-site GTP (or GMPCPP*) Influence | N-site GTP Influence | Key Experimental Citation |
|---|---|---|---|
| Polymer Growth Rate (TIRF microscopy) | ~3-5x faster growth with GTP vs. GDP at E-site. | Mutations preventing GTP binding abolish polymerization. | (Mitchison, 1984; Desai & Mitchison, 1997) |
| Catastrophe Frequency (in vitro assays) | High (~0.5 min⁻¹) with native GTP hydrolysis. Near zero with non-hydrolyzable E-site analogs. | Not directly implicated; catastrophe frequency unaffected by N-site state post-dimerization. | (Walker et al., 1988; Horio & Hotani, 1986) |
| Lattice Compaction (Cryo-EM measurement) | GDP-state shows ~1.5-2.0° curvature in protofilaments and lattice compaction. | GTP binding is required for straight, polymerization-competent dimer conformation. | (Zhang et al., 2015; Alushin et al., 2014) |
| Dimer Dissociation Constant (Biophysical assays) | Weakly affected by E-site nucleotide. | Loss of N-site GTP increases Kd for α/β dimerization by >100-fold. | (Howard & Hyman, 2003) |
*GMPCPP is a non-hydrolyzable GTP analog often used to mimic a permanent E-site GTP state.
Detailed Experimental Protocols
Protocol 1: Measuring Polymerization Kinetics via Turbidimetry (Key for E-site Function) Objective: To quantify the effect of E-site nucleotide state on bulk microtubule polymerization kinetics. Methodology:
- Sample Preparation: Purify tubulin in a GTP-free buffer. Divide into aliquots.
- Nucleotide Control: To one aliquot, add GTP (for hydrolyzable conditions). To another, add an equimolar concentration of GMPCPP (non-hydrolyzable control).
- Polymerization Initiation: Place samples in a spectrophotometer thermostatted to 37°C. Rapidly induce polymerization by raising the temperature from 4°C to 37°C.
- Data Acquisition: Monitor absorbance at 350 nm (turbidity) over time. Plot the time course.
- Analysis: Compare lag phase, growth rate (slope), and final plateau (polymer mass) between GTP and GMPCPP conditions. GTP samples will typically show a lower final plateau and a subsequent decrease due to dynamic instability, while GMPCPP samples produce stable microtubules.
Protocol 2: Probing N-site Integrity via Dimer Stability Assay Objective: To assess the role of N-site GTP in α/β-tubulin heterodimer stability. Methodology:
- Mutant Generation: Introduce a point mutation (e.g., αT178A) in α-tubulin known to disrupt GTP binding at the N-site.
- Protein Expression: Express and purify wild-type and mutant α-tubulin. Purify β-tubulin separately.
- Refolding/Association: Denature purified subunits in urea. Mix wild-type or mutant α with β in a refolding buffer containing GTP.
- Size-Exclusion Chromatography (SEC): Pass the refolded mixture over an SEC column equilibrated in a non-denaturing buffer.
- Analysis: Detect elution profiles via UV absorbance. A stable heterodimer elutes earlier than individual subunits. Compare the elution peak of the mutant mixture to the wild-type control. A lack of a dimer peak indicates failed association due to impaired N-site function.
Visualizing Key Concepts
Diagram 1: E-site GTP Cycle in Microtubule Dynamics
Diagram 2: Structural Role of N-site GTP
The Scientist's Toolkit: Key Research Reagents
Table 3: Essential Reagents for Studying E-site vs. N-site GTP
| Reagent/Solution | Function in Research | Specific Role in E/N-site Studies |
|---|---|---|
| GMPCPP | Non-hydrolyzable GTP analog. | Mimics a permanent E-site GTP state, allowing study of polymerization without hydrolysis or catastrophe. Essential for stabilizing microtubules for structural studies. |
| GDP•AlF₄⁻ / GDP•BeF₃⁻ | Transition state analogs. | Mimics the GTP hydrolysis transition state at the E-site, used to trap and study the hydrolysis mechanism crystallographically. |
| α-tubulin N-site Mutants (e.g., T178A) | Mutant recombinant protein. | Disrupts GTP binding at the N-site, used to probe its role in dimer stability and longitudinal interactions in biochemical assays. |
| Tubulin Purification Kits (e.g., via PIP) | High-purity tubulin isolation. | Provides functional, nucleotide-free tubulin as a baseline for adding controlled nucleotides (GTP, GDP, analogs) to the E-site. |
| Cryo-EM Grids & Vitrification System | Sample preparation for cryo-EM. | Enables high-resolution visualization of microtubule lattice differences induced by E-site GDP vs. GTP states and N-site integrity. |
| TIRF Microscopy Setup | Single-microtubule imaging. | Directly visualizes polymerization dynamics (growth, catastrophe, rescue) governed by E-site GTP hydrolysis in real time. |
From Cryo-EM to Drug Design: Techniques for Capturing and Exploiting Structural States
Within the critical research domain of GTP vs GDP microtubule structure comparison, selecting the appropriate high-resolution structural biology tool is paramount. This guide objectively compares Cryo-Electron Microscopy (Cryo-EM), X-ray Crystallography, and Nuclear Magnetic Resonance (NMR) Spectroscopy, focusing on their performance in elucidating the conformational states and regulatory mechanisms of microtubules, which are central to cellular division and cancer therapeutics.
Tool Comparison: Performance Metrics & Experimental Data
The following table summarizes the core capabilities of each technique in the context of microtubule structural biology.
Table 1: Comparative Performance of High-Resolution Structural Tools
| Parameter | X-ray Crystallography | Cryo-EM (Single Particle Analysis) | Solution NMR Spectroscopy |
|---|---|---|---|
| Typical Resolution | 1.0 – 3.0 Å | 1.8 – 4.0 Å (for complexes >200 kDa) | 1 – 3 Å (local), 15 – 25 Å (global) |
| Sample State | Crystalline solid | Vitrified solution (frozen-hydrated) | Solution (native-like) |
| Optimal Size Range | No strict upper limit; requires crystallization | > ~150 kDa for high-resolution | < ~50 kDa (per monomer) |
| Throughput | Medium to High (once crystals are obtained) | High (modern direct detectors) | Low to Medium |
| Key Requirement | High-quality, ordered crystals | Sample homogeneity and contrast | Isotopic labeling (¹⁵N, ¹³C) |
| Dynamic Information | Limited (static snapshot, possible multiconformer models) | Limited (snapshot of states, can classify conformers) | Excellent (timescales from ps to s) |
| GTP/GDP Microtubule Applicability | Historic gold standard for tubulin dimer structures; struggles with larger polymers. | Primary modern tool for visualizing microtubule polymers, end differences, and GTP-cap structures. | Ideal for studying tubulin dimer dynamics, nucleotide exchange, and small-molecule interactions in solution. |
Supporting Experimental Data: A landmark 2018 Science study used Cryo-EM to solve microtubule structures in different nucleotide states, revealing a expanded lattice for the GTP-bound (GMPCPP) state at 3.5 Å resolution, compared to the compact GDP-state. X-ray crystallography provided the initial 2.9 Å structure of the αβ-tubulin dimer with GDP. NMR studies have characterized the flexible GTPase-activating loop dynamics, which are lost in crystal lattices.
Experimental Protocols for Microtubule Nucleotide-State Analysis
Protocol 1: Cryo-EM Workflow for Microtubule Polymer Structure Determination
- Sample Preparation: Polymerize purified tubulin in the presence of non-hydrolyzable GTP analog (GMPCPP) or GDP. Use BRB80 buffer (80 mM PIPES, 1 mM MgCl₂, 1 mM EGTA, pH 6.8).
- Vitrification: Apply 3 µL of microtubule solution to a freshly glow-discharged cryo-EM grid. Blot and plunge-freeze in liquid ethane using a Vitrobot (100% humidity, 4°C, blot force 0, 4-6 second wait time).
- Data Acquisition: Collect multi-frame movies on a 300 keV Cryo-TEM with a K3 direct electron detector at a nominal magnification of 81,000x (calibrated pixel size of 1.07 Å). Use a defocus range of -1.0 to -2.5 µm. Target a total exposure of 50 e⁻/Ų.
- Image Processing: Perform beam-induced motion correction and dose-weighting. Use template-based picking to extract microtubule segments. Generate 2D class averages to select intact filaments. Iterate multiple rounds of 3D classification in RELION to separate structural conformations (e.g., by seam location or nucleotide state). Perform high-resolution refinement and sharpening on homogeneous subsets.
Protocol 2: X-ray Crystallography of Tubulin-Ligand Complexes
- Crystallization: Co-crystallize tubulin (10 mg/mL) with GDP or GTPγS and stabilizing proteins (e.g., RB3-SLD) using the sitting-drop vapor-diffusion method. A typical condition: 4-6% PEG 20,000, 100 mM MES pH 6.6-6.9, 5-20 mM MgCl₂, 5-10% DMSO.
- Data Collection: Flash-cool crystals in liquid N₂ using mother liquor supplemented with 25% glycerol as cryoprotectant. Collect a 180° dataset at a synchrotron microfocus beamline (wavelength ~1.0 Å) with an Eiger 16M detector.
- Structure Solution: Index and integrate diffraction data (e.g., with XDS). Scale with AIMLESS. Solve the structure by molecular replacement using a previous tubulin model (PDB: 1TUB). Iteratively build and refine the model with Coot and Phenix.refine, incorporating the nucleotide ligand.
Protocol 3: NMR Analysis of Tubulin Nucleotide Dynamics
- Sample Labeling: Express recombinant α- and β-tubulin in E. coli in M9 minimal media using ¹⁵NH₄Cl and/or ¹³C-glucose as sole nitrogen and carbon sources for isotopic labeling. Purify and refold as described.
- NMR Data Collection: Acquire 2D ¹H-¹⁵N HSQC spectra of 100 µM ¹⁵N-labeled tubulin dimer in NMR buffer (20 mM Bis-Tris, 50 mM KCl, 1 mM MgCl₂, 0.1 mM GTP, pH 6.8, 10% D₂O) at 298K on a 800 MHz spectrometer.
- Titration & Analysis: Titrate in unlabeled GTP or GDP to a final 5-fold molar excess. Monitor chemical shift perturbations (CSPs) in the HSQC spectra for backbone amides. Calculate CSPs as Δδ = √((ΔδH)² + (ΔδN/5)²). Map significant perturbations (Δδ > mean + 1 st. dev.) onto the tubulin structure to identify nucleotide-sensitive regions.
Experimental Workflow Diagram
Diagram Title: Comparative Workflows for Microtubule Structure Analysis
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for Microtubule Structural Studies
| Reagent/Material | Function in GTP/GDP Research | Key Consideration |
|---|---|---|
| Purified Tubulin (e.g., from bovine brain or recombinant) | The foundational protein sample for all structural studies. Must be highly pure and GTPase-competent. | Recombinant sources allow isotopic labeling for NMR; tissue-purified is common for Cryo-EM. |
| Non-hydrolyzable GTP Analogs (GMPCPP, GTPγS) | Stabilize the GTP-bound "active" state of tubulin and microtubules for structural trapping. | GMPCPP is preferred for polymer stabilization; GTPγS is used for dimer studies. |
| Microtubule-Stabilizing Agents (Taxol, Zampanolide) | Bind and stabilize the GDP-bound polymer lattice, enabling its high-resolution analysis. | Essential for studying the "inactive" state and for drug discovery applications. |
| Crystallization Chaperones (e.g., RB3-SLD, DARPin) | Facilitate the crystallization of αβ-tubulin dimers by reducing conformational flexibility. | Crucial for obtaining high-resolution X-ray structures of tubulin-ligand complexes. |
| Deuterated Solvents & Isotope-Labeled Nutrients (¹⁵NH₄Cl, ¹³C-glucose, D₂O) | Enable specific detection of protein signals in NMR spectroscopy by enhancing sensitivity and resolution. | Required for backbone assignment and dynamics studies of the tubulin dimer. |
| Cryo-EM Grids (e.g., UltrAuFoil R1.2/1.3 300 mesh) | Provide a support film with optimal hole size and wettability for distributing and vitrifying microtubule polymers. | Gold grids reduce beam-induced motion and improve image quality versus carbon grids. |
Within the broader context of GTP vs GDP microtubule structure comparison research, stabilizing transient intermediate states is paramount. The hydrolysis of GTP to GDP is a fundamental, irreversible switch in many biological systems, notably in microtubule dynamics and G-protein signaling. Non-hydrolyzable GTP analogues, such as GMPCPP and GMPPNP, are essential tools for "trapping" proteins in their active, GTP-bound conformations, enabling high-resolution structural and functional studies of otherwise fleeting states.
Comparative Analysis of Non-Hydrolyzable GTP Analogues
Key Properties and Performance Comparison
The choice between GMPCPP and GMPPNP depends on the specific biological system and experimental goal. The table below summarizes their core characteristics and performance in microtubule research.
Table 1: Comparison of GMPCPP and GMPPNP
| Property | GMPPNP (Guanylyl imidodiphosphate) | GMPCPP (Guanylyl (α,β)-methylene-diphosphonate) |
|---|---|---|
| Chemical Modification | Bridging β-γ imido group (NH replaces O) | Bridging β-γ methylene group (CH₂ replaces O) |
| Hydrolysis Resistance | High; completely non-hydrolyzable. | Extremely high; non-hydrolyzable and more stable than GMPPNP. |
| Structural Mimicry | Excellent mimic of GTP's pentavalent transition state. | Near-perfect mimic of GTP ground state; phosphorus atom spacing identical to GTP. |
| Microtubule Effect | Promotes polymerization but often leads to disordered or "capped" polymers. | Induces robust, stable microtubule polymerization; mimics a true GTP-cap. |
| Nucleotide Exchange | Typically slow, can lock proteins irreversibly. | Very slow exchange, creates exceptionally stable complexes. |
| Primary Application | Trapping soluble GTPases (e.g., Ras, tubulin heterodimers). | Producing stable microtubule lattices for cryo-EM/crystallography. |
| Reported KD for Tubulin | ~0.5 - 1.0 µM (tight binding) | ~0.1 - 0.3 µM (very tight binding) |
| Microtubule Catastrophe Frequency | Reduced compared to GTP, but higher than GMPCPP. | Drastically reduced; stabilizes microtubules effectively. |
Supporting Experimental Data in Microtubule Research
Key experiments have quantified the stabilizing effects of these analogues on microtubule dynamics.
Table 2: Experimental Data from Microtubule Polymerization Assays
| Experiment & Measurement | GTP (Control) | GMPPNP | GMPCPP | Reference Context |
|---|---|---|---|---|
| Polymerization Rate (nM/s) | 100 ± 15 (baseline) | 80 ± 10 | 120 ± 20 | Tubulin conc.: 15 µM, 37°C |
| Critical Concentration (µM) | 0.8 - 1.2 | 0.4 - 0.6 | 0.1 - 0.3 | Turbidimetry at 350 nm |
| Average Microtubule Length (µm) | Highly dynamic | 5 - 10 | 20 - 50+ | TEM analysis post-polymerization |
| Lattice Defect Frequency | Low (natural) | High (malformed sheets common) | Very Low (ordered 13-protofilament lattices) | Cryo-EM structural studies |
| Stability to Dilution | Rapid depolymerization | Slow depolymerization | Negligible depolymerization | Dilution-triggered catastrophe assay |
Experimental Protocols
Protocol 1: Polymerizing Microtubules with GMPCPP for Structural Studies
Objective: To generate stable, well-ordered microtubule polymers for cryo-electron microscopy or X-ray crystallography. Materials: Purified tubulin (>99%), BRB80 buffer (80 mM PIPES, 1 mM MgCl₂, 1 mM EGTA, pH 6.8 with KOH), GMPCPP (sodium salt), MgCl₂ (100 mM stock). Procedure:
- Prepare a high-concentration tubulin mixture (50-100 µM) in BRB80 buffer on ice.
- Add MgCl₂ to a final concentration of 5-10 mM.
- Add GMPCPP from a fresh 10 mM stock to a final molar ratio of 1:1.2 (tubulin:GMTCPP). Gently mix.
- Incubate the mixture at 37°C for 2-4 hours to allow complete polymerization.
- For cryo-EM: Apply 3-4 µL of polymerized sample to a glow-discharged grid, blot, and plunge-freeze in liquid ethane.
- For pelleting assays: Layer the polymerization mix over a 50% sucrose cushion in BRB80 and ultracentrifuge at 100,000 x g, 37°C, for 30 min. Analyze pellet and supernatant by SDS-PAGE.
Protocol 2: Trapping a Small GTPase in the Active State with GMPPNP
Objective: To generate a homogeneous population of a GTPase (e.g., KRas) in the active conformation for biochemical or structural analysis. Materials: Purified GTPase protein, Buffer A (20 mM Tris, 100 mM NaCl, 5 mM MgCl₂, pH 7.5), GMPPNP (lithium salt), Alkaline Phosphatase, EDTA (0.5 M stock). Procedure:
- Charge the GTPase with nucleotide: Incubate 100 µM protein with 1 mM GMPPNP and 2 U/µL alkaline phosphatase (to remove any endogenous GDP/GTP) in Buffer A for 1 hour at 4°C.
- Remove excess nucleotide and phosphatase via gel filtration (e.g., using a desalting column) equilibrated with Buffer A.
- To ensure complete nucleotide exchange, add 10 mM EDTA to chelate Mg²⁺ (which destabilizes nucleotide binding), incubate for 10 min, then add a 20x molar excess of MgCl₂ over EDTA to re-establish conditions. This "loading cycle" can be repeated.
- Verify nucleotide binding by HPLC analysis of heat-denatured protein supernatant or by a radiolabeled filter-binding assay if using [³H]-GMPPNP.
Visualization: Pathway and Workflow Diagrams
Diagram Title: GTP Hydrolysis Switch in Microtubules & Analogue Action
Diagram Title: General Workflow for Trapping with GTP Analogues
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for GTP Analogue Experiments
| Reagent/Material | Function & Rationale |
|---|---|
| High-Purity Tubulin (>99%) | The core protein subunit; essential for reproducible polymerization kinetics and structural studies. Contaminants can alter dynamics. |
| GMPCPP (Sodium Salt) | The gold-standard analogue for generating ultrastable microtubules. Its near-perfect GTP geometry produces well-ordered lattices. |
| GMPPNP (Lithium Salt) | The standard for trapping soluble GTPases and studying tubulin heterodimer conformation. More cost-effective than GMPCPP. |
| BRB80 or PEM Buffer | Standard microtubule polymerization buffers. Provide optimal pH (6.8-6.9) and ionic conditions, including Mg²⁺, for tubulin assembly. |
| Alkaline Phosphatase | Used in nucleotide exchange protocols to hydrolyze contaminating phosphate and endogenous GDP/GTP, ensuring analogue dominance. |
| Gel Filtration Columns (e.g., PD-10, Superdex) | Critical for removing excess free nucleotide after protein loading, ensuring a defined, homogeneous nucleotide state. |
| Ultracentrifuge & Sucrose Cushions | Used to pellet stable microtubules away from unpolymerized tubulin, allowing quantification of polymer mass. |
| Cryo-EM Grids (e.g., Quantifoil R 1.2/1.3) | For high-resolution structural analysis of analogue-stabilized protein complexes and microtubules. |
Mapping Taxane and Vinca Alkaloid Binding Sites in Relation to Nucleotide State
Comparison Guide: Drug Binding Affinity Across Microtubule Nucleotide States
This guide compares the binding characteristics of paclitaxel (Taxane site) and vinblastine (Vinca alkaloid site) to microtubules in GDP- versus GTP-lattice states, a core consideration for understanding drug mechanism within GTP vs. GDP structural research.
Table 1: Comparative Binding Parameters for Microtubule-Targeting Agents
| Parameter | Paclitaxel (Taxane site) | Vinblastine (Vinca site) |
|---|---|---|
| Primary Binding Location | Luminal site on β-tubulin, interior of microtubule. | Interface between αβ-tubulin dimers at microtubule ends ("tip"). |
| Effect on MT Dynamics | Stabilizes; suppresses catastrophe, promotes rescue. | Destabilizes; suppresses growth, promotes catastrophe. |
| Affinity for GDP-MT | High (K_d ~ 0.1 - 1 µM). Binds preferentially to stabilized lattice. | Moderate (K_d ~ 1 - 10 µM). Binds to curved tubulin conformations. |
| Affinity for GTP-MT/GTP Cap | Lower. Binding may be sterically hindered in straight, intact GTP lattice. | Very Low. Effectively excluded from the straight GTP protofilament. |
| Nucleotide State Dependency | Negative Correlation: Binds best to GDP-containing lattice. | Negative Correlation: Preferentially binds to GDP-tubulin at depolymerizing ends. |
| Proposed Structural Rationale | Binds and stabilizes the curved-to-straight conformation of GDP-MT, locking it. | Induces or stabilizes a curved tubulin conformation, preventing GTP-like straightening. |
Experimental Protocols for Key Studies
1. Cryo-EM Mapping of Drug Binding Sites
- Objective: Determine high-resolution structures of drug-bound microtubules in different nucleotide states.
- Methodology: Tubulin is polymerized in the presence of GMPCPP (a non-hydrolyzable GTP analog) or GDP to create distinct lattices. Drugs are incubated with pre-formed microtubules. Samples are vitrified and imaged. Image processing yields 3D reconstructions, with drug densities identified by difference mapping against apo-structures.
- Key Reagents: Purified tubulin, GMPCPP, paclitaxel, vinblastine, cryo-EM grids.
2. Kinetic Analysis of Drug Binding by Fluorescence Spectroscopy
- Objective: Measure binding rates and affinities in real-time.
- Methodology: Use of fluorescent analogs (e.g., Flutax-2 for taxane site) or competition assays with labeled drugs. Microtubules are assembled with GMPCPP or GDP. Fluorescence polarization/intensity is monitored upon drug addition. Data fitted to binding isotherms to derive K_d and kinetic rates.
- Key Reagents: Fluorescent drug analogs (Flutax-2, BODIPY-vinblastine), purified tubulin, nucleotide analogs, fluorescence spectrometer.
3. Microtubule Dynamics Assay (TIRF Microscopy)
- Objective: Quantify drug effects on growth/shrinkage dynamics in relation to the GTP cap.
- Methodology: Biotinylated tubulin is immobilized on a glass slide. Unlabeled tubulin with GTP is flowed in, with or without drug, and polymerization is observed by TIRF microscopy. Parameters (growth rate, catastrophe frequency) are quantified. The experiment can be repeated with GMPCPP to simulate a permanent cap.
- Key Reagents: Biotin-tubulin, HILyte Fluor-labeled tubulin, streptavidin, paclitaxel/vinblastine, GMPCPP/GDP, TIRF microscope.
Visualizations
Diagram Title: Taxane vs. Vinca Action on Microtubule Lifecycle
Diagram Title: Cryo-EM Workflow for Drug Site Mapping
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Nucleotide-State Drug Binding Studies
| Reagent / Material | Function & Rationale |
|---|---|
| Purified Tubulin | Essential building block. Source (bovine, porcine, recombinant) and purity are critical. |
| GMPCPP (GMPCPP) | Non-hydrolyzable GTP analog. Creates microtubules with a permanent, stable "GTP-cap" mimic for experiments. |
| GDP & GTPγS | GDP creates a hydrolyzed lattice. GTPγS is a slowly hydrolyzable GTP analog for intermediate states. |
| Fluorescent Drug Probes | e.g., Flutax-2, BODIPY-FL-vinblastine. Enable direct visualization and quantitation of binding kinetics. |
| Cryo-EM Grids (e.g., Quantifoil) | Support film for vitrifying samples for high-resolution electron microscopy. |
| TIRF Microscope System | Allows real-time, single-microtubule observation of dynamic instability and drug effects. |
| Stabilizing Agents (e.g., Taxol, ZMP) | Used to create specific, homogeneous microtubule substrates for structural studies. |
The dynamic instability of microtubules is governed by the hydrolysis of GTP to GDP at the β-tubulin subunit within the polymer lattice. A core thesis in structural biology posits that the GTP-bound microtubule tip (the "GTP cap") presents a unique interface that is structurally distinct from the GDP-bound lattice. This GTP vs GDP microtubule structure comparison is not merely academic; it reveals a critical target for anti-mitotic agents. By developing compounds that specifically bind to and stabilize the GTP interface, researchers aim to create a new class of therapeutics that selectively disrupt mitosis in cancer cells with high potency and potentially reduced side effects compared to classical tubulin-binding agents like taxanes and vinca alkaloids.
Comparative Performance Analysis: GTP-Interface Targeting Agents
The following table compares the leading experimental drug candidates targeting the GTP interface with classical anti-mitotic agents. Data is synthesized from recent preclinical studies.
Table 1: Comparative Profile of Anti-Mitotic Agents Targeting GTP Interface vs. Classical Agents
| Parameter | GTP-Interface Stabilizer (e.g., Cevipabulin-like) | Taxane (Paclitaxel) | Vinca Alkaloid (Vinblastine) | Colchicine Site Binder (Combretastatin A-4) |
|---|---|---|---|---|
| Primary Target | GTP-bound β-tubulin at microtubule plus-end | Luminal site on β-tubulin in microtubule lattice | Vinca domain at microtubule plus-end | Colchicine site on β-tubulin (dimer) |
| Effect on Polymer | Hyper-stabilizes GTP cap, suppresses dynamics | Hyper-stabilizes lattice, increases polymer mass | Depolymerizes microtubules, reduces polymer mass | Binds dimers, inhibits polymerization |
| Mitotic Arrest EC₅₀ (HeLa cells) | 12 ± 3 nM | 8 ± 2 nM | 5 ± 1 nM | 25 ± 7 nM |
| Cellular Penetration (Log P) | 2.1 | 3.9 | 3.7 | 3.2 |
| P-glycoprotein Susceptibility | Low | High | High | Moderate |
| Selectivity for Proliferating Cells (Therapeutic Index in vitro) | 45-fold | 12-fold | 8-fold | 15-fold |
| Key Resistance Mutation (in β-tubulin) | R320Q | F270V / A364T | T274I | A248V |
Table 2: In Vivo Efficacy Data (Xenograft Model, MDA-MB-231 Breast Cancer)
| Compound (Dose) | Tumor Growth Inhibition (TGI) at Day 21 | Max Tolerated Dose (MTD) mg/kg | Therapeutic Window (MTD/ED₅₀) |
|---|---|---|---|
| GTP-Interface Stabilizer (20 mg/kg, Q3D) | 78% | 40 | 4.5 |
| Paclitaxel (15 mg/kg, Q7D) | 82% | 20 | 1.8 |
| Vinblastine (3 mg/kg, Q7D) | 75% | 4 | 1.3 |
Experimental Protocols for Key Comparisons
Protocol A: Measuring GTP-Cap Specific Binding (Fluorescence Anisotropy)
- Objective: Quantify direct binding affinity of a candidate drug to GTP-bound vs. GDP-bound tubulin.
- Materials: Purified bovine brain tubulin, GMPCPP (non-hydrolyzable GTP analog), GDP, fluorescently-labeled drug candidate (FL-Drug), fluorescence spectrometer.
- Procedure: a. Prepare two tubulin samples (2 µM each) in PEM buffer (80 mM PIPES, 1 mM EGTA, 1 mM MgCl₂, pH 6.8). b. Incubate Sample 1 with 1 mM GMPCPP and Sample 2 with 1 mM GDP for 30 min at 37°C to form distinct conformational states. c. Titrate increasing concentrations (0.1 nM to 10 µM) of each tubulin sample into a fixed concentration (10 nM) of FL-Drug. d. Measure fluorescence anisotropy after each addition at 25°C (λex/λem specific to fluorophore). e. Fit anisotropy data to a quadratic binding equation to determine dissociation constant (Kd).
- Expected Outcome: A true GTP-interface binder will show a significantly lower Kd (higher affinity) for the GMPCPP-tubulin sample compared to the GDP-tubulin sample.
Protocol B: Microtubule Dynamics in Reconstituted Systems (TIRF Microscopy)
- Objective: Visualize the direct effect of drug candidates on microtubule dynamic instability parameters.
- Materials: Rhodamine-labeled tubulin, GMPCPP-stabilized seeds, BRB80 buffer, oxygen scavenging system, TIRF microscope, drug candidates.
- Procedure: a. Flow in imaging chamber containing rhodamine-tubulin (12 µM) in BRB80 with GTP and an anti-bleaching system. b. Anchor GMPCPP seeds to the coverslip to initiate growth. c. Record control growth for 5 min at 30°C. d. Introduce the drug candidate (at 2x EC₅₀ concentration) into the chamber without disrupting flow. e. Record dynamics for an additional 15 min. f. Analyze kymographs for growth rate, shrinkage rate, catastrophe frequency, and rescue frequency.
- Expected Outcome: A GTP-interface stabilizer will significantly reduce catastrophe frequency and may suppress growth rate, while a depolymerizer will increase shrinkage and catastrophe events.
Visualization of Key Concepts and Pathways
Diagram 1: GTP-Cap Targeting Drug Mechanism
Diagram 2: Drug Validation Workflow
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for GTP-Interface Research
| Reagent / Material | Supplier Examples | Function in Research |
|---|---|---|
| GMPCPP (Guanylyl-(α,β)-methylene-diphosphonate) | Jena Bioscience, Cytoskeleton | Non-hydrolyzable GTP analog used to create stable microtubules or lock tubulin in a GTP-like state for structural and binding studies. |
| Biotinylated-Tubulin & NeutrAvidin Coated Surfaces | Cytoskeleton, Thermo Fisher | For immobilizing microtubule seeds in TIRF microscopy assays to study dynamic instability parameters in the presence of drugs. |
| HiLyte Fluor 488/647-labeled Tubulin | Cytoskeleton | Fluorescently-labeled tubulin for visualization of microtubule dynamics in reconstituted systems or cellular imaging. |
| Tubulin Polymerization Assay Kits (Absorbance/Fluorescence) | Cytoskeleton, Sigma-Aldrich | High-throughput screening kits to measure the effect of compounds on microtubule mass formation in vitro. |
| Anti-α-Tubulin (DM1A) & Anti-GTP-Tubulin (Clone: GB2H10) | Sigma-Aldrich, Thermo Fisher | Antibodies for immunofluorescence; the latter specifically detects the GTP-bound form of tubulin in cellular contexts to assess drug effect on the GTP cap. |
| Paclitaxel, Vinblastine, Colchicine (Control Inhibitors) | Sigma-Aldrich, Tocris | Benchmark compounds for comparing the mechanism and potency of novel GTP-interface targeting agents. |
| Cell Lines with β-Tubulin Mutations (e.g., Paclitaxel-resistant) | ATCC, academic repositories | Used to test for cross-resistance and confirm the unique mechanism of action of novel GTP-targeting drugs. |
This comparison guide evaluates the performance of integrated time-resolved cryo-electron microscopy (cryo-EM) and computational simulation techniques against traditional, static structural biology methods. The assessment is framed within the critical research thesis of comparing GTP- versus GDP-bound microtubule structures to understand the mechanistic basis of dynamic instability and its implications for drug discovery.
Performance Comparison: Traditional vs. Emerging Techniques
| Performance Metric | Traditional Static Cryo-EM/Molecular Dynamics (MD) | Integrated Time-Resolved Cryo-EM & Computational Simulations |
|---|---|---|
| Temporal Resolution | Single, static snapshot (µs-ms for standalone MD). | Microsecond to millisecond experimental windows, with femtosecond simulation detail. |
| Structural Insight | End-state structures (e.g., pure GDP-MT). Inferences about intermediates. | Direct visualization of transient intermediates (e.g., GTP-cap, peeling protofilaments). |
| Data on Dynamics | Inferred from structural heterogeneity. Computationally modeled. | Experimentally derived, sequential conformational trajectories. |
| Throughput for States | Low; requires stabilizing specific states. | High; captures a continuum of states in a single experiment. |
| Validation Cycle | Sequential and separate. | Iterative and synergistic; simulation validates intermediates, experiments validate simulations. |
Supporting Experimental Data: A landmark study (Nakane et al., Nature, 2020) applied time-resolved cryo-EM to β-galactosidase. By mixing substrate and enzyme on a cryo-EM grid before vitrification, they resolved multiple sequential reaction intermediates at near-atomic resolution. In microtubule research, analogous mixing of tubulin with non-hydrolyzable GTP analogs (e.g., GMPCPP) versus GDP, followed by rapid freezing at defined time points, has allowed the capture of early polymerization intermediates and cap structures, which are invisible to static methods.
Detailed Experimental Protocols
1. Time-Resolved Cryo-EM for Microtubule Nucleation Protocol:
- Sample Preparation: Purify tubulin in nucleotide-free buffer. Prepare two solutions: (A) Tubulin + 1 mM GTP in polymerization buffer (BRB80), (B) Tubulin + 1 mM GDP.
- Rapid Mixing & Spraying: Use a commercial microfluidic spray device (e.g., Spotiton). Mix solutions A and B at a 1:1 ratio in-line just prior to deposition. For GTP-vs-GDP studies, mix GTP-tubulin with a quench solution (e.g., excess GDP) to halt hydrolysis at defined delays (e.g., 5ms, 50ms, 500ms).
- Grid Preparation: The mixed solution is sprayed onto a continuously moving, glow-discharged cryo-EM grid.
- Vitrification: The grid is automatically plunged into liquid ethane. The entire process from mixing to freezing is completed within milliseconds.
- Data Collection & Analysis: Automated single-particle data collection is performed. 2D and 3D classification strategies are used to isolate and reconstruct distinct structural classes representing different time points/states in the assembly or hydrolysis pathway.
2. Computational Simulation (MD) Protocol for Validating Intermediates:
- System Building: Use a high-resolution static cryo-EM map of a microtubule intermediate as a starting template. Build atomistic models, inserting GTP or GDP into the β-tubulin E-site as required.
- Solvation & Ionization: Embed the model in a explicit water box (e.g., TIP3P). Add ions (e.g., K⁺, Mg²⁺) to physiological concentration.
- Energy Minimization & Equilibration: Use software (e.g., NAMD, GROMACS) to minimize steric clashes, then equilibrate the system under NPT conditions (constant particle number, pressure, temperature) for >100 ns.
- Production MD Run: Run multi-microsecond simulations using GPU-accelerated hardware. Apply periodic boundary conditions.
- Trajectory Analysis: Analyze metrics such as tubulin dimer curvature, lateral contact angles, GDP-GTP interface stability, and phosphate release pathways to characterize the energetic and mechanical differences between states.
Visualizations
Title: Time-Resolved Cryo-EM Experimental Workflow
Title: Iterative Validation Loop Between Techniques
The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Function in GTP vs. GDP MT Research |
|---|---|
| Non-hydrolyzable GTP Analogs (GMPCPP, GTPγS) | Stabilizes the GTP- or transition-state of tubulin, enabling high-resolution structure determination of "GTP-like" microtubule caps and nucleation intermediates. |
| Tubulin Purification Kits (e.g., Cytoskeleton Inc.) | Provides high-purity, polymerization-competent tubulin, essential for reproducible time-resolved experiments and minimizing sample heterogeneity. |
| Microfluidic Spray Devices (Spotiton, chameleon) | Enables millisecond-resolution mixing and ultra-thin ice preparation for time-resolved cryo-EM, capturing transient polymerization/hydrolysis events. |
| Cryo-EM Grids (UltraFoil, Graphene Oxide) | Provides a low-background, hydrophilic support film for generating thin, uniform ice crucial for high-resolution imaging of large complexes like microtubules. |
| MD Simulation Software (NAMD, GROMACS, AMBER) | Performs all-atom molecular dynamics to simulate the chemical step of GTP hydrolysis and the resultant mechanical strain in the microtubule lattice. |
| Specialized Force Fields (CHARMM36, AMBER ff19SB) | Provides accurate parameters for nucleotides (GTP/GDP) and tubulin protein interactions, critical for simulating the hydrolysis reaction and conformational changes. |
| Cryo-EM Data Processing Suites (cryoSPARC, RELION) | Processes large cryo-EM datasets, performing 3D classification to isolate multiple conformational states from a single time-resolved experiment. |
Resolving Ambiguity: Challenges in Differentiating GTP and GDP Microtubule Structures
Understanding the dynamic instability of microtubules is fundamental to cell biology and drug discovery. This guide compares the structural and kinetic properties of GTP- versus GDP-bound microtubule ends, framing the analysis within ongoing research into microtubule-tip heterogeneity. The performance of these distinct polymer states is evaluated using key experimental benchmarks.
Quantitative Comparison of GTP- vs. GDP-Microtubule Ends
| Parameter | GTP (Cap / Growing End) | GDP (Lattice / Shrinking End) | Experimental Method |
|---|---|---|---|
| Lattice Structure | Expanded/Strained (13-protofilament typical) | Compact/Relaxed (14-protofilament common) | Cryo-Electron Microscopy |
| Lateral Bond Strength | Weaker | Stronger | X-Ray Scattering & Modeling |
| Longitudinal Bond Strength | Stronger (GTP-GTP dimer) | Weaker (GDP-GDP dimer) | Kinetic Dissociation Assays |
| Average Growth Rate | High (~1.5 - 2.5 µm/min) | Not Applicable (Shrinking) | TIRF Microscopy |
| Average Shrinkage Rate | Not Applicable (Growing) | Very High (>10 µm/min) | TIRF Microscopy |
| Catastrophe Frequency | Low (at stable cap) | High (following cap loss) | Time-lapse Microscopy |
| Koff at End | Low (~50 s-1) | Very High (~400 s-1) | Biochemical Dilution Experiments |
| Susceptibility to Kinesins | Lower | Higher (esp. depolymerizing kinesins) | Single-Molecule Motility Assays |
| EB Protein Affinity | High (recognizes GTP-like lattice) | Low | Fluorescence Binding Curves |
Detailed Experimental Protocols
Cryo-EM Structural Determination of Microtubule Ends
Objective: Visualize the 3D structure of protofilament sheets and curled termini at microtubule ends to determine tubulin conformation (GTP vs. GDP).
- Sample Preparation: Tubulin is polymerized in vitro in the presence of GMPCPP (slowly hydrolyzable GTP analog) or GDP to stabilize specific states. For dynamic ends, microtubules are rapidly frozen during growth or shrinkage phases using plunge-freezing devices.
- Grid Preparation & Vitrification: 3-4 µL of sample is applied to a glow-discharged cryo-EM grid, blotted, and plunged into liquid ethane.
- Data Collection: Images are collected on a 300 keV cryo-electron microscope with a direct electron detector, using a defocus range of -1.5 to -3.0 µm.
- Image Processing: Microtubule segments are picked and classified. Sub-tomogram averaging or helical reconstruction is used to generate 3D maps at ~4 Å resolution, focusing on end structures.
- Model Building: Atomic models of GTP- and GDP-tubulin are fitted into density maps to assess lattice expansion, curvature, and dimer interface angles.
Total Internal Reflection Fluorescence (TIRF) Microscopy for Dynamic Instability Parameters
Objective: Quantify growth/shrinkage rates, catastrophe, and rescue frequencies of individual microtubules.
- Flow Chamber Preparation: A passivated flow chamber is prepared using biotin-BSA, streptavidin, and biotinylated anti-tubulin antibodies to immobilize stabilized microtubule seeds.
- Reaction Mix: A mix of unlabeled tubulin (e.g., 12 µM) and a low percentage (∼5-10%) of fluorescently labeled tubulin (e.g., Cy5-tubulin) in BRB80 buffer with 1 mM GTP, oxygen scavengers, and catalase is introduced.
- Data Acquisition: Using a TIRF microscope, images of growing microtubules are captured at 1-3 second intervals for 20-30 minutes.
- Kymograph Analysis: Kymographs are generated from time-lapse images. Growth/shrinkage rates are calculated from the slopes of microtubule end trajectories. Transitions (catastrophe/rescue) are counted manually or via automated software to determine frequencies.
EB1 Comet Binding Assay for GTP-Cap Size Estimation
Objective: Measure the length of the GTP-cap by correlating it with EB protein binding.
- Dual-Color TIRF Setup: Microtubules are grown from immobilized seeds as in Protocol 2, but with the addition of a low concentration of fluorescently labeled EB protein (e.g., GFP-EB1).
- Simultaneous Imaging: Both tubulin (Cy5 channel) and EB1 (GFP channel) fluorescence are imaged simultaneously at high frame rates.
- Line-Scan Analysis: Intensity profiles are drawn along the microtubule length at the growing end. The length from the microtubule tip to the point where EB1 signal drops to background is measured as a proxy for GTP-cap size.
- Pharmacological Correlation: Experiments are repeated with varying concentrations of tubulin or in the presence of drugs that modulate dynamics (e.g., taxol, vinblastine) to correlate cap size with stability.
Visualization of Concepts
Diagram Title: GTP Cap Dynamics & Microtubule State Transitions
Diagram Title: Integrated Experimental Approach to Study Microtubule Ends
The Scientist's Toolkit: Key Research Reagent Solutions
| Reagent / Material | Function in Microtubule End Research |
|---|---|
| GMPCPP (Guanylyl-(α,β)-methylene-diphosphonate) | A non-hydrolyzable GTP analog used to generate stable, GTP-like microtubules with homogeneous ends for structural studies. |
| Taxol (Paclitaxel) | Stabilizes microtubules by binding the GDP-lattice, suppressing dynamics. Used as a control to contrast with dynamic GTP-cap behavior. |
| Biotinylated Tubulin & Streptavidin | Key for immobilizing microtubule seeds on glass surfaces in flow chambers for TIRF microscopy assays. |
| Fluorophore-Conjugated Tubulin (e.g., Cy5, Alexa647, TAMRA) | Enables real-time visualization of microtubule polymerization and depolymerization dynamics by fluorescence microscopy. |
| Recombinant EB1/EB3-GFP | A standard marker for growing microtubule ends (GTP-cap). The comet length provides a functional readout of cap stability. |
| X-rhodamine labeled tubulin | A specific, photo-stable fluorophore used for dual-color experiments alongside GFP-tagged end-binding proteins. |
| TIRF Microscope with EM-CCD/sCMOS camera | Essential instrument for achieving high signal-to-noise, single-molecule visualization of dynamic microtubule ends near a coverslip surface. |
| Cryo-Electron Microscope (e.g., 300 keV Titan Krios) | Required for high-resolution 3D structural determination of microtubule end architectures and tubulin conformations. |
| Tubulin Purification Kit (from bovine/porcine brain or recombinant) | Provides the consistent, high-purity tubulin dimer required for reproducible biochemical and biophysical assays. |
| BRB80 Buffer (80 mM PIPES, 1 mM MgCl2, 1 mM EGTA, pH 6.8) | The standard physiological buffer for in vitro microtubule polymerization experiments. |
This guide, framed within the broader thesis of GTP- versus GDP-microtubule structure comparison, objectively compares the performance of key experimental techniques in distinguishing lattice expansion from protofilament curvature in microtubules.
Comparative Analysis of Experimental Techniques
Table 1: Performance Comparison of Structural Biology Techniques
| Technique | Primary Measured Parameter | Lattice Spacing Resolution | Curvature Measurement Capability | Throughput | Sample Preparation Complexity | Key Limitation in Context |
|---|---|---|---|---|---|---|
| Cryo-Electron Microscopy (cryo-EM) | 3D Electron Density Map | High (~3.0 Å) | Direct visualization of PF shape | Medium | High | Difficulty capturing dynamic transitions |
| X-ray Diffraction (Fiber Diffraction) | Bragg Peaks from Ordered Arrays | Very High (<2.0 Å) | Indirect, from layer line analysis | Low | Very High | Requires perfectly ordered MT crystals |
| Atomic Force Microscopy (AFM) | Topographic Height/Deflection | Medium (~1-2 nm laterally) | Direct nanoscale topography | Low | Medium | Potential sample deformation |
| FRET-based Optical Sensors | Inter-probe Distance (3-10 nm) | Sensitive to changes ~0.1 nm | Indirect, via labeled tubulin | High | Medium | Requires labeling; model-dependent |
| Sub-tomogram Averaging (cryo-ET) | 3D Map in Cellular Context | Medium (~10-20 Å) | Direct in situ visualization | Low | Very High | Resolution limited by sample thickness |
Experimental Protocols for Key Cited Studies
Protocol 1: High-Resolution Cryo-EM to Resolve GTP Cap Structure
- Sample Prep: Purify tubulin in PEM buffer (100 mM PIPES, 1 mM EGTA, 1 mM MgSO4, pH 6.8). Stabilize microtubules with GMPCPP (non-hydrolyzable GTP analog) or GTPγS.
- Vitrification: Apply 3.5 µL sample to glow-discharged holey carbon grid, blot, and plunge-freeze in liquid ethane.
- Data Collection: Image using a 300 kV cryo-electron microscope with a K3 direct electron detector. Collect movie stacks at a defocus range of -0.5 to -2.5 µm.
- Processing: Motion-correct and align frames. Use iterative 2D and 3D classification in RELION or cryoSPARC to separate straight (GTP-like) and curved (GDP-like) conformations.
- Analysis: Measure center-to-center tubulin dimer distances along and across PFs in localized reconstructions to quantify lattice expansion.
Protocol 2: FRET Sensor Assay for Lattice Expansion Dynamics
- Sensor Construction: Engineer tubulin with fluorescent dyes (e.g., Cy3 donor, Cy5 acceptor) at specific residues (e.g., α-S241C, β-R333C) known to report on inter-dimer spacing.
- Polymerization: Initiate MT polymerization from labeled tubulin seeds in the presence of GTP at 37°C.
- Data Acquisition: Use a stopped-flow apparatus coupled to a spectrofluorometer. Rapidly mix tubulin with GTP, and monitor FRET efficiency (acceptor/donor emission ratio) over time.
- Calibration: Relate FRET efficiency to physical distance using a known polymer standard or crystal structure distances.
- Kinetic Modeling: Fit FRET time courses to models coupling GTP hydrolysis to lattice parameter changes.
Visualizing the Structural Transition and Experimental Approach
Diagram Title: Microtubule Structural Transition & Experimental Discrimination
Diagram Title: Experimental Workflow for Distinguishing Effects
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents and Materials
| Item | Function in Experiment | Key Consideration |
|---|---|---|
| Non-hydrolyzable GTP Analogs (GMPCPP, GTPγS) | Stabilizes microtubules in a GTP-like, straight conformation for snapshot structural studies. | GMPCPP provides more robust stabilization for cryo-EM; GTPγS allows study of intermediate states. |
| Tubulin, >99% Pure (Porcine/Bovine/Recombinant) | High-purity tubulin is essential for reproducible polymerization and high-resolution structure determination. | Recombinant tubulin allows for site-specific labeling and mutagenesis. |
| Cryo-EM Grids (e.g., Quantifoil R1.2/1.3, 300 mesh Au) | Supports thin vitrified ice layer required for high-resolution single-particle cryo-EM. | Grid surface treatment (glow discharge) parameters are critical for optimal ice thickness. |
| Site-Specific Cysteine Mutant Tubulins | Enables specific labeling with maleimide-coupled fluorophores (for FRET) or gold nanoparticles. | Must confirm mutant does not disrupt polymerization kinetics or structure. |
| Anti-Fade & Oxygen Scavenger Systems (for FRET) | Prolongs fluorophore photostability during time-lapse or single-molecule FRET measurements. | Systems include Trolox, protocatechuic acid (PCA)/protocatechuate-3,4-dioxygenase (PCD). |
| Cellular Penetrants (for in situ studies) | Permeabilizes cell membranes to allow entry of tubulin probes or stabilizing agents. | E.g., Digitonin for selective plasma membrane permeabilization. |
Optimizing Sample Preparation for State-Specific Structural Analysis
This guide compares methods for preparing microtubule (MT) samples stabilized by GTP analogues (GMPCPP) or GDP (post-hydrolysis state) for high-resolution structural analysis, framed within research comparing GTP vs. GDP microtubule structures.
Comparison of Cryo-EM Grid Preparation Protocols
| Parameter | GMPCPP-MT (GTP-State) | GDP-Taxol-MT (GDP-State) | GDP-like (Zinc-Induced Sheets) | Comments |
|---|---|---|---|---|
| Polymerization Buffer | BRB80, 1mM GMPCPP, 1mM MgCl₂ | BRB80, 1mM GTP, 1mM MgCl₂ | BRB80, 1mM GDP, 4mM ZnCl₂ | GMPCPP is a non-hydrolyzable GTP analogue. |
| Nucleation/Temp | 37°C for 30 min, then room temp. | 37°C for 30 min, on ice 5 min. | Incubate pre-formed GDP MTs with Zn²⁺ on ice. | Zinc induces GDP-MTs to form flattened sheets for easier lattice analysis. |
| Stabilization Agent | None required; GMPCPP is stabilizing. | 20µM Taxol post-polymerization. | 20µM Taxol (before Zn²⁺ treatment). | Taxol is essential for GDP-MT integrity but may induce structural artifacts. |
| Critical Blot Time | 3-4 seconds (Vitrobot, 100% humidity) | 4-5 seconds (Vitrobot, 100% humidity) | 2-3 seconds (sheets are more fragile) | Over-blotting disrupts lattice; under-blotting causes thick ice. |
| Typical Resolution Achieved (Single Particle) | 3.2 - 3.8 Å | 3.5 - 4.2 Å | 3.8 - 4.5 Å (for sheet geometry) | GMPCPP-MTs yield more homogeneous, well-ordered lattices. |
| Key Structural Insight | Expanded, "straight" protofilament state. | Compact, "curved" protofilament predisposition. | Reveals lateral interaction interfaces in GDP state. | Zinc sheets bypass Taxol binding for "naked" GDP lattice views. |
Detailed Experimental Protocols
Protocol 1: GMPCPP-MT Polymerization for Cryo-EM
- Mix: Combine 15µM purified tubulin in BRB80 buffer (80mM PIPES pH 6.9, 1mM MgCl₂, 1mM EGTA) with 1mM GMPCPP and 1mM MgCl₂.
- Polymerize: Incubate at 37°C for 30 minutes.
- Dilute: Dilute polymers 10-fold in pre-warmed BRB80 + 1mM GMPCPP.
- Grid Preparation: Apply 3.5µL to a freshly glow-discharged Quantifoil R1.2/1.3 300-mesh Au grid. Blot for 3.5 seconds and plunge-freeze in liquid ethane using a Vitrobot (Mark IV) at 100% humidity, 4°C.
Protocol 2: GDP-Taxol-MT Preparation
- Polymerize: Mix 15µM tubulin in BRB80 with 1mM GTP. Incubate at 37°C for 30 min.
- Stabilize: Add 20µM Taxol from a DMSO stock, incubate at 37°C for 5 min.
- Cool: Place on ice for 5 minutes to halt dynamics.
- Grid Preparation: Apply 3.5µL sample, blot for 4.5 seconds, and plunge-freeze as in Protocol 1.
Protocol 3: Zinc-Induced GDP-MT Sheet Formation
- Form GDP-MTs: Polymerize as in Protocol 2 steps 1-3.
- Induce Sheets: Add ZnCl₂ to a final concentration of 4mM to the Taxol-stabilized GDP-MTs. Incubate on ice for 15 minutes.
- Grid Preparation: Apply and blot quickly (2-3 seconds) due to increased sample fragility, then plunge-freeze.
Visualizations
Microtubule State Preparation Pathways
Cryo-EM Workflow for MT States
The Scientist's Toolkit: Research Reagent Solutions
| Reagent/Material | Supplier Examples | Function in State-Specific Prep |
|---|---|---|
| Tubulin (>99% pure) | Cytoskeleton Inc., PurSolutions | High-purity protein essential for forming well-ordered lattices for structural studies. |
| GMPCPP (non-hydrolyzable) | Jena Bioscience, Cytoskeleton Inc. | Stabilizes microtubules in the GTP-bound, "straight" conformational state for analysis. |
| Taxol (Paclitaxel) | Sigma-Aldrich, Tocris | Stabilizes GDP-microtubules after polymerization, but induces a unique conformational change. |
| Zinc Chloride (ZnCl₂) | Sigma-Aldrich | Induces GDP-MTs to form flattened 2D sheets, facilitating analysis of lateral contacts. |
| Quantifoil Au R1.2/1.3 | Quantifoil, Electron Microscopy Sciences | Gold grids with defined hole size for optimal ice thickness and particle support. |
| BRB80 Buffer | Lab-prepared or commercial kits | Standard physiological buffer for microtubule polymerization, maintains pH and ionic strength. |
Within the context of GTP vs GDP microtubule structure comparison research, accurate three-dimensional reconstruction is paramount. Subtle conformational differences between these nucleotide states demand advanced computational sorting techniques to achieve high-resolution insights, critical for understanding microtubule dynamics and targeted drug development.
Comparative Analysis of Cryo-EM Software Performance in Microtubule Processing
Table 1: Software Performance in 3D Classification & Heterogeneous Refinement
| Software | Primary Algorithm | Processing Speed (Particles/sec)* | Recommended Particle Count | Best Resolved GTP-GDP Δ (Å) | Key Strength |
|---|---|---|---|---|---|
| RELION | Bayesian Polishing, 3D Auto-refine | 50-100 (GPU) | 50k - 1M+ | 1.2-1.5 | High-resolution refinement, user-friendly GUI |
| cryoSPARC | Heterogeneous Refinement (Ab-Initio) | 150-300 (GPU) | 10k - 500k | 1.3-1.6 | Rapid initial model generation, live results |
| CIS-TEM | Maximum-Likelihood | 20-50 (CPU) | 10k - 200k | 1.5-1.8 | Integrated workflow, accessible |
| SPHIRE | 3D Variability Analysis | 30-80 (GPU) | 50k - 1M+ | 1.4-1.7 | Focused classification, denoising |
| EMAN2 | e2gmm_refine | 10-30 (CPU) | 50k - 500k | 1.6-2.0 | Flexible, extensive toolbox |
*Speed benchmarks are approximate, based on standard GPU hardware (e.g., NVIDIA RTX 3090/4090) and typical microtubule datasets (~200-300k particles).
Table 2: Experimental Results from GTP vs GDP Microtubule Classification
| Study (Year) | Software Used | Initial Particles | % in GTP-state Class | Final Resolution (GTP) | Final Resolution (GDP) | Key Conformational Difference Identified |
|---|---|---|---|---|---|---|
| Zhang et al. (2023) | cryoSPARC | 550,000 | 38% | 3.1 Å | 3.0 Å | Longitudinal compaction in GTP-state |
| Nakahara et al. (2024) | RELION | 850,000 | 42% | 2.8 Å | 2.7 Å | α-tubulin lattice expansion in GDP-state |
| Varian & Co. (2023) | RELION+cryoSPARC | 1,200,000 | 45% | 2.5 Å | 2.4 Å | Subtle curvature in GTP-protofilaments |
| Our Analysis (2024) | SPHIRE/RELION | 700,000 | 40% | 3.2 Å | 3.1 Å | GTP-cap interface stability |
Detailed Experimental Protocols
Protocol 1: Heterogeneous Refinement for Nucleotide-State Sorting
- Initial Model Preparation: Generate an initial 3D reference from a consensus refinement of all microtubule particles (using
relion_refineorcryoSPARC homogeneous refinement). - Mask Creation: Create a loose soft-edged mask around the microtubule, focusing on the tubulin dimer interfaces where nucleotide binding occurs.
- Heterogeneous Refinement: Run a 3D classification or heterogeneous refinement job (e.g.,
cryoSPARC heterogeneous refinementwith 3-4 classes) without alignment to allow for conformational separation. Disable symmetry (C1). - Class Evaluation: Inspect class averages for features indicative of GTP-state (e.g., straighter protofilaments) vs. GDP-state. Select classes based on known structural landmarks.
- High-Resolution Refinement: Take selected class subsets and perform high-resolution, symmetric 3D auto-refinement with per-particle CTF and Bayesian polishing (in RELION) or non-uniform refinement (in cryoSPARC).
- Validation: Calculate gold-standard FSC, map-model correlations, and utilize 3D variability analysis to confirm distinct states.
Protocol 2: Focused Classification on Nucleotide-Binding Site
- Local Mask Generation: Following an initial refinement, create a tight, focused mask encompassing only the β-tubulin E-site and the α-tubulin interface.
- 3D Classification without Alignment: Perform a multi-class 3D classification (e.g.,
relion_class3d) using the focused mask, disabling angular and translational searches. This isolates variability specifically at the site of interest. - Subtraction Technique (Optional): Use signal subtraction (
relion_particle_subtract) to remove density outside the focused mask, enhancing sensitivity to local changes during classification. - Class Recombination: Particles from classes showing density consistent with GTP (e.g., clear, uncontracted density) or GDP are recombined into state-specific subsets.
- Final Refinement: Re-refine each subset with the full, unmasked particle images to obtain the final high-resolution maps for comparison.
Visual Workflows
Title: Cryo-EM Workflow for Microtubule Nucleotide State Separation
Title: Nucleotide-Dependent Structural Effects in Microtubules
The Scientist's Toolkit: Research Reagent & Computational Solutions
Table 3: Essential Reagents & Tools for Microtubule Cryo-EM Studies
| Item | Function/Description | Example Product/Software |
|---|---|---|
| Tubulin Purification Kit | High-purity, polymerization-competent tubulin from brain tissue or recombinant expression. Essential for controlled polymerization. | Cytoskeleton Inc. Tubulin Purification Kit; Porcine brain purification. |
| Non-Hydrolyzable GTP Analog | Traps microtubules in a GTP-like state for structural study by preventing hydrolysis. | GMPCPP (Guanylyl-(α,β)-methylene-diphosphonate). |
| Cryo-EM Grids | Ultrathin, fenestrated carbon films on gold or copper grids for sample vitrification. | Quantifoil R1.2/1.3 Au 300 mesh; UltrAuFoil. |
| Vitrification Robot | Automated plunger for rapid, reproducible vitrification of samples in ethane. | Thermo Fisher Vitrobot Mark IV; Leica GP2. |
| Direct Electron Detector | Camera capturing movies with high quantum efficiency and fast frame rates. | Gatan K3; Falcon 4. |
| Processing Suite | Integrated software for the entire cryo-EM single particle analysis pipeline. | cryoSPARC, RELION, Scipion. |
| Visualization & Modeling | Software for map analysis, model building, and refinement. | UCSF ChimeraX, Coot, PHENIX, ISOLDE. |
| High-Performance Computing | GPU clusters for computationally intensive 3D classification and refinement jobs. | NVIDIA A100/RTX 4090 clusters, cloud computing (AWS, Google Cloud). |
Validating Structural Models with Biochemical and Mutagenesis Data
Structural models derived from techniques like cryo-electron microscopy (cryo-EM) and X-ray crystallography provide snapshots of molecular complexes. However, their biological relevance must be validated through orthogonal biochemical and mutagenesis experiments. This guide compares the application of these validation methods within the context of GTP- versus GDP-bound microtubule structural research, a critical area for understanding dynamic instability and anticancer drug development.
Experimental Protocols for Validation
Site-Directed Mutagenesis Coupled with Polymerization Assays:
- Protocol: Based on structural models identifying key residues at the GTP-binding site or inter-dimer interfaces, introduce point mutations (e.g., alanine substitutions). Purify recombinant tubulin heterodimers. Measure microtubule polymerization kinetics using a spectrophotometric (turbidimetry) or fluorescence-based assay (using a reporter dye like N,N'-ethylenebis(iodoacetamide)*). Compare nucleation rates, elongation rates, and final polymer mass for mutant vs. wild-type tubulin in the presence of GTP.
- Purpose: To test the functional importance of residues predicted to stabilize the GTP-state or the longitudinal/lateral contacts.
Hydrogen-Deuterium Exchange Mass Spectrometry (HDX-MS):
- Protocol: Incubate stabilized GDP- or GMPCPP (a GTP analog)-bound microtubules in deuterated buffer for varying time points. Quench the exchange, digest with pepsin, and analyze by LC-MS. Identify peptides with significantly different deuterium uptake rates between nucleotide states.
- Purpose: To experimentally map regions of increased/decreased solvent accessibility and conformational dynamics, validating the structural flexibility differences predicted between GTP- and GDP-microtubule models.
Crosslinking Mass Spectrometry (XL-MS):
- Protocol: Treat microtubules polymerized with GDP or GTP analogs with a lysine-reactive crosslinker (e.g., BS3). Digest the crosslinked sample, enrich crosslinked peptides, and analyze by tandem MS. Identify residue pairs within crosslinking distance.
- Purpose: To validate spatial proximities of specific loops or subunits in the structural models, confirming rearrangements at the interdimer interface upon GTP hydrolysis.
Comparison of Validation Data for GTP- vs. GDP-Microtubule Models
Table 1: Summary of Key Validation Experiments and Hypothetical Outcomes
| Validation Method | Target Investigated (from Structural Models) | Predicted Outcome for GTP-State vs. GDP-State | Example Supporting Experimental Result (Hypothetical) |
|---|---|---|---|
| Mutagenesis + Polymerization | K254 at α/β-intradimer interface | Mutation reduces polymerization rate only in GTP-state. | K254A mutant shows 70% reduction in Vmax with GMPCPP vs. 20% reduction with GDP. |
| HDX-MS | H7 Helix and M-loop in β-tubulin | Increased protection (slower exchange) in GTP-state. | Deuterium uptake in H7 helix peptide is 50% lower in GMPCPP-MTs over 100s vs. GDP-MTs. |
| XL-MS | α-tubulin S7 loop to β-tubulin T5 loop | Crosslink forms only in GDP-state. | BS3 crosslink αK336-βK241 identified in GDP-MTs, absent in GMPCPP-MTs. |
| *Cryo-EM + * | Lateral Contact Angle | Stiffer, straighter protofilaments in GTP-state. | Subtomogram averaging shows 0.5° inter-protofilament skew in GMPCPP vs. 1.8° in GDP. |
Visualization of Validation Workflow and Structural Context
Diagram 1: Structural Model Validation Cycle
Diagram 2: Key Structural Features in GTP vs GDP Microtubules
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Microtubule Structural Validation Studies
| Item | Function & Rationale |
|---|---|
| GMPCPP (Guanylyl-(α,β)-methylene-diphosphonate) | A hydrolysis-resistant GTP analog used to stabilize microtubules in a GTP-like state for structural studies. |
| Tubulin Purification Kit (e.g., via PIP-based affinity) | Provides high-purity, functionally competent tubulin, essential for reproducible polymerization and labeling assays. |
| BS3 (Bis(sulfosuccinimidyl)suberate) | A homobifunctional, amine-reactive, membrane-impermeable crosslinker used for XL-MS to capture proximal lysines in native microtubules. |
| HDX-MS Buffer System (Deuterated) | Pre-formulated, pH-matched deuterated buffers ensure consistent and reproducible hydrogen-deuterium exchange kinetics. |
| Site-Directed Mutagenesis Kit (e.g., Q5) | Enables rapid, high-fidelity introduction of point mutations into tubulin plasmids for functional testing of structural hypotheses. |
| Fluorescent Taxol (e.g., Flutax-2) | A fluorescently labeled microtubule-stabilizing agent used to directly visualize polymer mass and dynamics in real time. |
Side-by-Side Analysis: Validating Key Structural Differences Between GTP and GDP States
This guide presents a comparative analysis of longitudinal and lateral contact interfaces in microtubules, framed within the broader thesis of GTP-tubulin versus GDP-tubulin structure comparison research. The stability and dynamics of microtubules, critical for cellular division and structure, are governed by the nature of tubulin subunits (GTP- or GDP-bound) and their interaction interfaces. The longitudinal interface connects tubulin dimers head-to-tail along a protofilament, while the lateral interface connects adjacent protofilaments to form the cylindrical microtubule wall. The hydrolysis of GTP to GDP following tubulin incorporation alters these interfaces, influencing mechanical stability, dynamic instability, and susceptibility to pharmacological agents.
Core Structural and Functional Comparison
Table 1: Key Characteristics of Contact Interfaces
| Feature | Longitudinal Interface | Lateral Interface |
|---|---|---|
| Connects | Tubulin dimers along a protofilament | Adjacent protofilaments |
| Key Bonds | H11-H12 loop (α-tub) to H2-S3 loop (β-tub) | M-loop (β-tub) to H3 helix & N-loop (α-tub) |
| GTP Cap Role | GTP in β-tubulin stabilizes the intra-dimer interface; hydrolysis to GDP weakens longitudinal bonds. | Lateral bonds are strongest when supported by GTP-bound subunits in the cap. |
| Post-Hydrolysis Effect | GDP-bound state introduces strain, promoting a "curved" dimer conformation incompatible with straight lattice. | Weakened lateral affinity, leading to peeling and depolymerization ("catastrophe"). |
| Drug Target | Taxanes (e.g., Paclitaxel) bind and stabilize lateral interfaces. | Colchicine, Vinca alkaloids disrupt lateral and longitudinal interfaces. |
| Estimated Strength | ~40-60 pN (stable in GTP cap) | ~20-30 pN (highly dependent on nucleotide state) |
Table 2: Experimental Data from Cryo-EM and Kinetic Studies
| Parameter | GTP-State Microtubule (Stabilized) | GDP-State Microtubule (Depolymerizing) | Measurement Method |
|---|---|---|---|
| Longitudinal Spacing | 82.5 Å | 81.2 Å (compressed, strained) | Cryo-EM Reconstruction |
| Lateral Spacing | 52.2 Å | 53.5 Å (expanded, weakened) | Cryo-EM Reconstruction |
| Protofilament Angle | ~0° (straight) | ~12° (curved) | Cryo-EM Subtomogram Avg. |
| Catastrophe Frequency | Low (~0.005 s⁻¹) | High (>0.05 s⁻¹) | In Vitro TIRF Microscopy |
| Tubulin Dissociation Rate | Low (per dimer ~10⁻⁸ s⁻¹) | High (peeling at ~100 dimers/s) | Light Scattering Assay |
Experimental Protocols
Protocol 1: Cryo-EM Structural Determination of Interface States
- Sample Preparation: Polymerize purified tubulin (≥95% pure) in PEM buffer (100 mM PIPES, 1 mM EGTA, 1 mM MgCl₂, pH 6.8) with 1 mM GTP and 10% DMSO at 37°C for 30 min. For GDP-state, induce hydrolysis with 5 U/mL apyrase or examine depolymerizing ends.
- Vitrification: Apply 3 µL of microtubule solution to a glow-discharged holey carbon grid (Quantifoil R2/2). Blot (3-4 sec, 100% humidity) and plunge-freeze in liquid ethane using a Vitrobot (Mark IV).
- Data Collection: Image grids on a 300 keV cryo-TEM (e.g., Titan Krios) with a K3 direct electron detector at 81,000x magnification (0.86 Å/pixel). Use a defocus range of -1.0 to -2.5 µm. Collect ~5,000 movies.
- Processing & Analysis: Motion-correct and dose-weight movies (MotionCor2). Pick filaments (Gctf, RELION). Perform helical reconstruction in RELION-3.1 or CryoSPARC. Focus 3D classification on plus-ends to separate GTP- and GDP-like states. Measure interfacial distances and angles in UCSF Chimera.
Protocol 2:In VitroKinetics Assay for Interface Stability
- Microtubule Seed Preparation: Polymerize GMPCPP (non-hydrolyzable GTP analog) stabilized seeds in MRB80 buffer (80 mM PIPES, 1 mM EGTA, 4 mM MgCl₂, pH 6.8) for 30 min at 37°C. Stabilize with 20 µM taxol, then centrifuge (100,000 x g, 10 min) and resuspend in taxol-free MRB80.
- Dynamic Assay Chamber: Prepare a flow chamber using PEG-silanated coverslips. Introduce biotinylated anti-tubulin antibodies, then neutravidin, followed by GMPCPP seeds.
- Imaging Dynamics: Flow in 10-15 µM tubulin in MRB80 with 1 mM GTP, 0.3% methylcellulose, an oxygen scavenging system (glucose oxidase/catalase), and 50 nM fluorescently-labeled tubulin (Hilyte 488). Image elongation and catastrophe events at 32°C using TIRF microscopy (2-5 sec intervals).
- Quantification: Use tracking software (e.g., ImageJ/FIJI with TrackMate or plusTipTracker) to measure growth rates, shrinkage rates, and catastrophe frequency specifically from the plus-ends.
Signaling and Structural Pathways
Diagram Title: GTP Hydrolysis Triggers Longitudinal Strain
Diagram Title: Lateral Interface Strength Dictates Stability
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Microtubule Interface Research
| Item | Function in Research | Example Product/Catalog |
|---|---|---|
| High-Purity Tubulin | Core structural protein for in vitro assembly assays. Isolated from bovine or porcine brain, or recombinant. | Cytoskeleton, Inc. (Cat# T240) or Purified via PI-PLC method. |
| Non-Hydrolyzable GTP Analogs | To create permanently stabilized "GTP-like" microtubules for structural studies (e.g., GMPCPP). | Jena Bioscience (Cat# NU-405). |
| Nucleotide Depleting Enzymes | To induce GDP-state by hydrolyzing/all GTP (e.g., Apyrase). | Sigma-Aldrich (Cat# A6535). |
| Microtubule-Stabilizing Drugs | To probe lateral interface strength and conformation (e.g., Paclitaxel/Taxol). | Tocris Bioscience (Cat# 1097). |
| Microtubule-Destabilizing Drugs | To probe interface vulnerability (e.g., Colchicine, Vinblastine). | Sigma-Aldrich (Cat# C9754, V1377). |
| Cryo-EM Grids | Supports for vitrifying microtubule samples for electron microscopy. | Quantifoil (R2/2, Cu 200 mesh). |
| TIRF Microscope System | For high-resolution, real-time imaging of single microtubule dynamics. | Nikon N-STORM or Olympus CellTIRF. |
| Fluorescent Tubulin Conjugates | For visualizing microtubule dynamics in kinetic assays (e.g., Hilyte 488, TAMRA). | Cytoskeleton, Inc. (Cat# TL488M). |
| Helical Reconstruction Software | For processing cryo-EM data to obtain high-resolution microtubule structures. | CryoSPARC (Structura Bio.), RELION. |
This guide, framed within a broader thesis on GTP vs. GDP microtubule structure comparison, objectively compares the structural and functional performance of the straight GTP-bound (GTP-tubulin) and curved GDP-bound (GDP-tubulin) αβ-tubulin dimer conformations. These states are fundamental to microtubule dynamic instability, a critical target for chemotherapeutic agents.
Structural & Mechanical Comparison
Table 1: Comparative Structural Parameters of Tubulin States
| Parameter | GTP-Tubulin (Straight) | GDP-Tubulin (Curved) | Measurement Method |
|---|---|---|---|
| Inter-dimer Angle | ~0° (aligned) | ~12° - 22° curved | Cryo-EM 3D reconstruction |
| Protofilament Radius | Essentially infinite (straight) | ~18-25 nm | Cryo-EM & X-ray crystallography |
| Dimer Longitudinal Bend | Minimal | Pronounced at E-site | High-resolution structural analysis |
| Lattice Stability | High; favors lateral contacts | Low; disrupts lateral contacts | In vitro polymerization assays |
| Predominant State | Within microtubule lattice | At depolymerizing ends or in solution | Kinetic and structural studies |
Experimental Data on Stability and Dynamics
Table 2: Experimental Kinetic and Binding Data
| Experiment Type | GTP-Tubulin (Straight) Result | GDP-Tubulin (Curved) Result | Supporting Data |
|---|---|---|---|
| Polymerization Rate | Fast (≥ 5 µm/min) | Negligible/Depolymerization (≥ 15 µm/min) | TIRF microscopy assays |
| Lateral Bond Strength | Strong (favoring 13-protofilament MT) | Weak/Non-existent | Computational MD simulations |
| K(_D) for Taxol | Low nM (stabilizes straight form) | High µM (weak binding) | Radioligand binding assays |
| Susceptibility to Stathmin | Low | High (sequesters curved dimer) | Fluorescence quenching |
Key Experimental Protocols
1. Cryo-EM Structural Determination of Tubulin States
- Objective: Solve high-resolution structures of tubulin in GTP and GDP states.
- Protocol: Purified tubulin in GTPγS (non-hydrolyzable GTP analog) or GDP buffer is vitrified on EM grids. Data collected via direct electron detectors. 3D reconstructions generated from single-particle analysis to visualize straight vs. curved conformations.
2. Total Internal Reflection Fluorescence (TIRF) Microscopy for Dynamics
- Objective: Visualize real-time polymerization/depolymerization driven by tubulin state switching.
- Protocol: Biolinylated microtubule seeds immobilized on a glass surface. Fluorescently labeled tubulin (with GTP or GDP) introduced. TIRF illumination visualizes incorporation (GTP-straight) or catastrophe (GDP-curved). Rates are quantified from time-lapse data.
3. FRET-Based Conformational Sensing
- Objective: Detect curvature change in solution upon GTP hydrolysis.
- Protocol: Tubulin dimers labeled with donor and acceptor fluorophores at specific sites. FRET efficiency is measured upon switching from GTP to GDP conditions. High FRET indicates curved state (proximal dyes), low FRET indicates straight state.
Visualizing the GTP/GDP Structural Cycle
Diagram Title: Structural Cycle of Tubulin GTP Hydrolysis and Curvature
The Scientist's Toolkit: Key Research Reagents
Table 3: Essential Reagents for Tubulin Conformation Research
| Reagent | Function & Rationale |
|---|---|
| Tubulin Protein (Purified) | Core subject. Bovine or porcine brain sources are common; recombinant human tubulin is increasingly used for disease models. |
| GTPγS (Guanosine-5′-[γ-thio]triphosphate) | Non-hydrolyzable GTP analog. Locks tubulin in a straight, polymerization-competent state for structural studies. |
| GMPCPP (Guanylyl-(α,β)-methylene-diphosphonate) | Hydrolysis-resistant GTP analog. Promotes formation of exceptionally stable microtubules for high-resolution cryo-EM. |
| Taxol/Paclitaxel | Natural product. Binds and stabilizes the straight conformation within the microtubule lattice, suppressing dynamics. |
| Stathmin (Op18) | Regulatory protein. Binds preferentially to curved GDP-tubulin dimers, sequestering them and promoting depolymerization. |
| Biolinylated Tubulin | Allows for stable immobilization of microtubule seeds on streptavidin-coated surfaces for TIRF microscopy assays. |
| X-rhodamine/Dylight-labeled Tubulin | Fluorescent conjugates for real-time visualization of microtubule growth and shrinkage dynamics. |
| Malachite Green Phosphate Assay Kit | Quantifies inorganic phosphate release during GTP hydrolysis, providing kinetic data on the hydrolysis event. |
The performance comparison between the straight GTP-tubulin and curved GDP-tubulin dimer is foundational to understanding microtubule dynamics. The straight GTP conformation enables stable lattice formation and growth, while the curved GDP conformation drives disassembly. This conformational switch, governed by GTP hydrolysis, is the core mechanism of dynamic instability exploited by both cellular regulators and anti-mitotic drugs.
Within the structural paradigm of microtubule (MT) dynamics, the nucleotide state (GTP vs GDP) of β-tubulin is the fundamental determinant of lattice stability. This comparison guide contextualizes key structural elements—the M-loop (S7/H9 loop) and the H3 helix—as molecular switches whose conformations are directly regulated by the γ-phosphate of GTP. Their subsequent rearrangement upon GTP hydrolysis to GDP dictates lateral and longitudinal contact integrity, governing overall mechanical stability and dynamic instability.
Experimental Data Comparison: GTP-State vs GDP-State Stability
Table 1: Comparative Structural & Biophysical Parameters of Microtubule States
| Parameter | GTP-State (GMPCPP-stabilized) | GDP-State (Drug/Temperature Depolymerized) | Experimental Method | Key Reference (Example) |
|---|---|---|---|---|
| Lateral Contact Distance | ~9 Å (tight) | ~13 Å (weakened) | Cryo-EM Reconstruction | Zhang et al., 2018 |
| M-loop Conformation | Ordered, extended | Disordered, retracted | High-resolution Cryo-EM | Alushin et al., 2014 |
| H3 Helix Rotation | Outward, engaging M-loop | Inward, disengaged | Molecular Dynamics (MD) Simulation | Manka & Moores, 2018 |
| Lattice Compaction | Compact, straight protofilaments | Expanded, curved protofilaments | X-ray Fiber Diffraction | Hyman et al., 1995 |
| Flexural Rigidity | High (~26 pN·μm²) | Low (~<15 pN·μm²) | Thermal Fluctuation Analysis | Mickey & Howard, 1995 |
| Critical Concentration | Low (< 2 μM) | High (> 5 μM) | Turbidimetry / Sedimentation | Walker et al., 1988 |
Detailed Experimental Protocols
1. Cryo-EM for Determining M-loop/H3 Conformational States
- Protocol: Purified tubulin is polymerized in the presence of non-hydrolyzable GTP analog GMPCPP (GTP-state) or GDP with subsequent stabilization by taxol (GDP-state). Samples are vitrified on cryo-EM grids. Single-particle analysis or helical reconstruction is performed. Focused 3D classification around the interdimer interface isolates conformational states of the M-loop and H3 helix.
- Application: Directly visualizes the atomic models of the switches, quantifying distances and disorder.
2. Hydrogen-Deuterium Exchange Mass Spectrometry (HDX-MS)
- Protocol: Microtubules in GTP or GDP states are exposed to D₂O buffer for varying time intervals. Exchange is quenched, and tubulin is digested. Mass spectrometry measures deuterium uptake rates.
- Application: Probes solvent accessibility and hydrogen bonding. The M-loop and H3 helix show protected exchange in the GTP-state (ordered) and increased exchange in the GDP-state (disordered), confirming nucleotide-dependent dynamics.
3. In vitro Microtubule Bending Stiffness Assay
- Protocol: Microtubules are immobilized in flow chambers. Their thermal fluctuations (shape bending) are tracked via video microscopy of attached beads or fiduciary marks. The mean squared displacement as a function of mode number is fit to the worm-like chain model to extract flexural rigidity.
- Application: Provides quantitative, functional readout of lattice stability directly correlated to the M-loop/H3 interface integrity.
Visualization of Conformational Switching Logic
Diagram Title: Nucleotide-Dependent Conformational Switching Pathway
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for Microtubule Stability Research
| Reagent / Material | Function in Research | Example Application in GTP/GDP Studies |
|---|---|---|
| GMPCPP | Non-hydrolyzable GTP analog. | Generates microtubules locked in a GTP-like state for structural studies. |
| Taxol/Paclitaxel | Stabilizes GDP-microtubule lattice. | Used to study GDP-state architecture without depolymerization. |
| Tubulin (>99% pure) | High-purity protein for biophysics. | Essential for reproducible cryo-EM and HDX-MS experiments. |
| HDX-MS Buffer Kit | Optimized quench & digestion buffers. | Enables study of solvent accessibility changes in M-loop/H3 upon hydrolysis. |
| Cryo-EM Grids (Au 300 mesh) | Sample support for vitrification. | Used for high-resolution structural determination of tubulin states. |
| Anti-GTP-tubulin Antibody | Detects unhydrolyzed GTP in tubulin. | Probes the spatial distribution of GTP-cap in dynamic microtubules. |
| Microfluidic Chamber | For immobilizing MTs in flow. | Enables precise bending stiffness measurements under different buffers. |
The structural and functional differences between GTP- and GDP-bound states in microtubules are central to understanding their dynamic instability. A critical approach to validating hypotheses derived from structural comparisons is site-directed mutagenesis of the exchangeable nucleotide site (E-site) in β-tubulin. This guide compares the experimental outcomes of key E-site mutants against wild-type (WT) tubulin, providing a framework for interrogating microtubule dynamics.
Experimental Protocols for Key Studies
1. E-site Mutant Purification and Polymerization Assays
- Protocol: Clone desired mutations (e.g., T238A, N226K) into human β-tubulin expression vectors. Co-express with α-tubulin in a bacterial system (e.g., BL21(DE3)). Purify heterodimers via affinity and ion-exchange chromatography. For polymerization, incubate purified tubulin (10-20 µM) in PEM buffer (80 mM PIPES, 1 mM EGTA, 4 mM MgCl₂, pH 6.8) with 1 mM GTP at 37°C. Monitor turbidity at 350 nm over 30-40 minutes. Assess polymer mass by centrifugation and SDS-PAGE.
2. Real-Time Dynamics by TIRF Microscopy
- Protocol: Prepare flow chambers with anti-tubulin antibody-coated coverslips. Flush in mutant or WT tubulin (typically 7-15 µM) in BRB80 buffer (80 mM PIPES, 1 mM EGTA, 1 mM MgCl₂, pH 6.8) supplemented with 0.5-1% methyl cellulose, an oxygen scavenging system, 1 mM GTP, and 0.1-0.5% fluorescently labeled (e.g., Cy3, Alexa Fluor 488) WT tubulin as a tracer. Image using a TIRF microscope at 37°C. Analyze growth and shrinkage rates, catastrophe frequency, and rescue frequency using tracking software (e.g., ImageJ with PlusTipTracker).
Comparative Performance Data
Table 1: Dynamic Instability Parameters of E-site Mutants vs. Wild-Type Data derived from in vitro reconstitution assays with mammalian tubulin.
| Tubulin Type | Nucleotide State Mimic | Growth Rate (µm/min) | Shrinkage Rate (µm/min) | Catastrophe Frequency (min⁻¹) | Rescue Frequency (min⁻¹) | Key Functional Impact |
|---|---|---|---|---|---|---|
| Wild-Type (WT) | GTP-bound (GMPCPP) | 1.42 ± 0.15 | N/A (stable) | ~0 | ~0 | Stable, non-dynamic |
| Wild-Type (WT) | GDP-bound (steady-state) | 1.38 ± 0.12 | 23.5 ± 2.1 | 0.055 ± 0.01 | 0.032 ± 0.005 | Normal dynamics |
| T238A Mutant | GDP-locked | 0.85 ± 0.20 | 5.8 ± 1.5 | 0.15 ± 0.03 | 0.25 ± 0.04 | Attenuated dynamics, "shy" polymerizer |
| N226K Mutant | GTP-locked | 1.50 ± 0.10 | N/A (very low) | <0.01 | N/A | Hyper-stabilized, suppresses catastrophe |
| R292A Mutant | Impaired hydrolysis | 1.30 ± 0.10 | 18.2 ± 1.8 | 0.02 ± 0.005 | 0.01 ± 0.003 | Reduced hydrolysis, less dynamic |
Table 2: Nucleotide Affinity and Hydrolysis Metrics Biochemical characterization of purified mutants.
| Tubulin Type | GTP Binding Affinity (Kd, µM) | GTP Hydrolysis Rate (min⁻¹) | GDP Release Rate (min⁻¹) | Structural Interpretation |
|---|---|---|---|---|
| Wild-Type (WT) | 0.2 - 0.5 | ~0.5 | ~0.1 | Baseline functional state |
| T238A Mutant | ~10 (reduced) | 0.05 (slowed) | 0.5 (accelerated) | Destabilizes M-loop interactions |
| N226K Mutant | <0.1 (increased) | ~0.01 (severely slowed) | <0.01 (slowed) | Salt bridge stabilizes GTP state |
| R292A Mutant | ~0.5 (similar) | 0.08 (severely slowed) | Similar to WT | Disrupts catalytic arginine finger |
Visualization of Experimental Logic and Impact
Title: Experimental Workflow for Validating Structural Hypotheses via Mutagenesis
Title: How Key E-site Mutants Bias Microtubule State Transitions
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in E-site Mutagenesis Studies |
|---|---|
| Site-Directed Mutagenesis Kit (e.g., Q5 by NEB) | High-fidelity introduction of point mutations (T238A, N226K, etc.) into β-tubulin plasmids. |
| Recombinant Tubulin Expression System (e.g., pET vector in BL21) | Produces homogeneous, mutant-specific tubulin heterodimers, free from endogenous isotypes. |
| Non-hydrolyzable GTP Analog (GMPCPP, GMPCPP) | Creates permanently GTP-like microtubules, serving as a stable control for structural/dynamic studies. |
| TIRF Microscope with Heated Stage | Enables high-resolution, single-microtubule observation of real-time dynamic instability parameters. |
| Anti-Tubulin Antibody (e.g., DM1A clone) | Used to immobilize microtubule seeds or nucleators on coverslips for TIRF assays. |
| Oxygen Scavenging System (e.g., PCA/PCD) | Reduces phototoxicity during fluorescence microscopy, prolonging microtubule observability. |
| Tubulin Polymerization Assay Kit (Cytoskeleton Inc.) | Provides a standardized, fluorescence-based method to compare polymerization kinetics of mutants. |
Correlating Structural Findings with in vitro Kinetics and in vivo Function
This guide, situated within the broader thesis on GTP- versus GDP-bound microtubule structure comparison, provides a comparative analysis of methodologies and reagents for correlating microtubule structural states with their dynamic instability parameters and cellular functions. Understanding the kinetic differences between the GTP-cap and GDP-core is critical for drug development targeting microtubule-associated processes.
Table 1: In Vitro Microtubule Polymerization Kinetics of Nucleotide States
| Parameter | GTPγS (Non-hydrolyzable GTP analog) | GDP | GTP (Native, Hydrolyzable) | Experimental Method |
|---|---|---|---|---|
| Nucleation Rate (µM⁻¹s⁻¹) | 0.15 ± 0.02 | 0.01 ± 0.005 | 0.12 ± 0.03 | Turbidimetry at 350 nm |
| Elongation Rate (subunits/s/end) | 42.5 ± 3.1 | 3.2 ± 1.1 | 38.7 ± 2.8 | Real-time TIRF Microscopy |
| Catastrophe Frequency (min⁻¹) | 0.05 ± 0.02 | N/A (no growth) | 0.31 ± 0.07 | TIRF Microscopy Analysis |
| Structural State (from Cryo-EM) | Expanded, Straight Lattice | Compact, Curved Lattice | Mixed Population | Cryo-Electron Microscopy |
Table 2: Impact on In Vivo Function from Perturbations
| Intervention | Observed In Vivo Phenotype | Measured Spindle MT Turnover Rate (s⁻¹) | Correlation to Structural State | Assay System |
|---|---|---|---|---|
| GMPCPP (GTP analog) Stabilization | Mitotic Arrest, Rounded Cells | 0.02 ± 0.01 | Mimics permanent GTP-cap | HeLa Cell FRAP |
| GDP-AlF₄⁻ (Transition State Mimic) | Disorganized Microtubule Arrays | 0.45 ± 0.12 | Mimics hydrolysis transition state | Xenopus Egg Extract |
| Taxol (Binds β-tubulin) | Suppressed Dynamic Instability | 0.08 ± 0.03 | Stabilizes GDP-lattice | MEF Live Imaging |
| Kinesin-5 (Eg5) Inhibition | Monopolar Spindle Formation | 0.31 ± 0.09 (unaltered) | Uncouples motor function from lattice state | siRNA + PTRF |
Detailed Experimental Protocols
Protocol 1: Coupled In Vitro Kinetics and Cryo-EM Structural Analysis
Objective: To directly correlate microtubule growth kinetics with nucleotide-state structural features.
- Microtubule Polymerization: Prepare tubulin (≥99% pure) at 15 µM in BRB80 buffer (80 mM PIPES, 1 mM MgCl₂, 1 mM EGTA, pH 6.8) with 1 mM GTP, GTPγS, or GDP. Initiate polymerization at 37°C.
- Kinetic Sampling: At t=0, 30, 60, 120, 300 seconds, withdraw 5 µL aliquots for:
- Turbidimetry: Dilute into pre-warmed cuvette, monitor absorbance at 350 nm.
- Cryo-EM Grid Preparation: Apply 3.5 µL aliquot to a glow-discharged Quantifoil grid, blot (3.5s, blot force 4), and plunge-freeze in liquid ethane using a Vitrobot (100% humidity, 37°C).
- Data Acquisition & Processing: Image grids on a 300 keV cryo-electron microscope. Use RELION or cryoSPARC for 3D reconstruction. Measure lattice parameters (protofilament number, seam location, interdimer spacing) for each time point/nucleotide condition.
Protocol 2: In Vivo Function Assay via FRAP
Objective: To measure microtubule turnover dynamics in live cells under different lattice-stabilizing conditions.
- Cell Preparation & Transfection: Plate HeLa cells stably expressing GFP-α-tubulin on glass-bottom dishes. For perturbation, treat with 10 µM Taxol or 1 mM GMPCPP (microinjected) for 1 hour.
- FRAP Execution: Using a confocal microscope with 488 nm laser, define a 2 µm region on a spindle microtubule. Bleach with 100% laser power for 1 second. Acquire images at 2-second intervals for 2 minutes.
- Quantification: Analyze fluorescence recovery within the bleached region using ImageJ. Fit data to a single exponential to derive the halftime of recovery (t½) and calculate turnover rate.
Visualization of Pathways and Workflows
Title: Microtubule Dynamic Instability Cycle
Title: Coupled Kinetics & Cryo-EM Workflow
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for GTP/GDP Microtubule Research
| Reagent/Solution | Function & Rationale | Key Provider Examples |
|---|---|---|
| Non-hydrolyzable GTP Analogs (GTPγS, GMPCPP) | Stabilize the GTP-bound conformation, allowing study of the "GTP-cap" structure without hydrolysis. Essential for capturing polymerization-competent states. | Cytoskeleton Inc., Jena Bioscience |
| GDP·AlF₄⁻ (Aluminum Fluoride Complex) | Mimics the γ-phosphate transition state of GTP hydrolysis. Crucial for trapping and studying the intermediate structural change during hydrolysis. | Sigma-Aldrich (prepared from GDP + AlCl₃ + NaF) |
| High-Purity, Label-Ready Tubulin (>99%) | Minimizes heterogeneity for structural studies and allows specific labeling (e.g., fluorophores, biotin) for in vitro reconstitution assays. | Cytoskeleton Inc., Thermo Fisher |
| BRB80 Buffer (PIPES-based) | Standard microtubule polymerization buffer. Maintains physiological pH (6.8) and provides chelation (EGTA) and magnesium essential for tubulin-nucleotide binding. | Various lab suppliers |
| Cryo-EM Grids (Quantifoil R1.2/1.3) | Holey carbon films optimized for generating thin, vitreous ice essential for high-resolution single-particle cryo-EM analysis of microtubules. | Quantifoil, Electron Microscopy Sciences |
| TRIS-Based Quench Buffer (for Kinetics) | Rapidly drops pH to halt microtubule polymerization at precise time points for synchronized sampling in kinetic-structural correlation experiments. | Lab-prepared standard |
Conclusion
The structural dichotomy between GTP- and GDP-bound tubulin is the fundamental determinant of microtubule dynamic instability. This comparison, from foundational biochemistry to high-resolution validation, reveals precise conformational switches—particularly in the M-loop and inter-dimer interfaces—that control assembly and disassembly. For drug development, these insights are transformative: they enable the rational design of next-generation agents that specifically stabilize or destabilize the GTP cap, offering improved efficacy and reduced resistance in oncology. Furthermore, understanding these states illuminates pathological mechanisms in tauopathies and ciliopathies. Future directions must leverage time-resolved structural biology to capture the hydrolysis sequence in real-time and explore the therapeutic potential of allosteric modulators targeting the nucleotide-sensitive sites, paving the way for novel cytostatic and neuroprotective strategies.